Integrated Genome-Wide Association Study Findings: Identification of a Neurodevelopmental Network for Attention Deficit Hyperactivity Disorder
Abstract
Objective:
Attention deficit hyperactivity disorder (ADHD) is a highly heritable neuropsychiatric disorder. In the present study, the authors investigated the presence of genomic convergence in the top findings of the five published genome-wide association studies (GWASs) of ADHD.
Method:
The authors carried out bioinformatics pathway analyses, using the Ingenuity and BiNGO tools, as well as a systematic literature analysis of 85 genes from the five published GWASs containing single nucleotide polymorphisms associated with ADHD at a p value <0.0001.
Results:
Findings revealed that 45 of the 85 top-ranked ADHD candidate genes encode proteins that fit into a neurodevelopmental network involved in directed neurite outgrowth. Data on copy number variations in patients with ADHD and data from animal studies provide further support for the involvement of this network in ADHD etiology. Several network proteins are also directly modulated by stimulants, the most commonly used psychopharmacological treatment for ADHD.
Conclusions:
The authors have identified a protein network for ADHD that contributes to our understanding of the molecular basis of the disorder. In addition, the data suggest new candidate genes for ADHD and provide clues to future research into psychopharmacological ADHD treatments.
Attention deficit hyperactivity disorder (ADHD) is a common neuropsychiatric disorder that is observed in children and adults. ADHD is characterized by age-inappropriate levels of hyperactive, impulsive, and inattentive behaviors and leads to significant clinical and psychosocial impairments (DSM-IV-TR). The disorder has a worldwide prevalence of 5%–6% in children (1) and 1%–4% in adults (2, 3). The most effective and commonly used medications for ADHD are the stimulants methylphenidate and amphetamine and the nonstimulant drug atomoxetine, which (predominantly) modulate dopaminergic and noradrenergic neurotransmission (4, 5).
Converging neurobiological evidence suggests that ADHD involves alterations of catecholaminergic brain circuits (6). Neuroimaging studies in children with ADHD also show neurodevelopmental brain anomalies, such as reduced cortical volume and folding and gray matter heterotopia (7–10), which indicate that, neuroanatomically, ADHD is a disorder of early brain development. In addition, specific neuroimaging modalities, such as diffusion tensor imaging (11–13), (resting-state) functional magnetic resonance imaging (fMRI) (14–16), and EEG (17), have shown that both structural and functional brain connectivity are impaired in ADHD patients.
Evidence from family and twin studies shows that ADHD is a highly heritable disorder, and approximately 76% of the phenotypic variance can be explained by genetic factors (18). ADHD behaves as a multifactorial (or complex) disorder, in which different combinations of genetic and environmental factors contribute to the overall risk of developing the disorder. The genetic model underlying most cases of ADHD is likely one in which multiple genetic factors of small-to-moderate effect size contribute to the disease etiology (19). Recent data indicate that rare copy number variations of genes may also be relevant to the etiology of ADHD (20–22), but pending further studies, it is unclear which proportion of ADHD cases could be explained by these variants.
To date, eight independent genome-wide linkage scans have been conducted for ADHD (23, 24). A recent meta-analysis of seven of these studies identified genome-wide significant linkage for a chromosomal region on 16q (23). In addition, a large number of candidate gene association studies have been published, and these have primarily focused on genes involved in catecholaminergic neurotransmission. Recent meta-analyses of these studies have yielded significant meta-association for six ADHD candidate genes (i.e., the dopamine and serotonin transporter genes DAT1/SLC6A3 and 5-HTT/SLC6A4, the DRD4 and DRD5 dopamine receptor genes, the HTR1B serotonin receptor gene, and the SNAP25 gene involved in neurotransmission) (25, 26). In 2008, the results of three genome-wide association studies (GWASs) for ADHD were published (27–29) in two independent data sets. Lasky-Su et al. (28) reported the following two findings that reached (trait-specific) genome-wide significance: 1) a single nucleotide polymorphism (SNP) in CDH13 and 2) a SNP in GFOD1.
Very recently, the results of two additional GWASs for ADHD in two independent samples were reported (30, 31). Apart from CDH13, which maps to the only chromosomal region that yielded genome-wide significant linkage for ADHD in a recent meta-analysis (23), overlap between GWAS findings and any of the reported linkage or candidate gene-based association findings has been limited (32), although this is probably due in part to the fact that the published GWASs for ADHD are strongly underpowered.
In the present study, we attempted to integrate the most significant findings from the five reported GWASs for ADHD into protein signaling networks that would not only increase our understanding of the genetic etiology of ADHD but also provide clues for future research into more effective (psychopharmacological) treatments for the disorder. Based on bioinformatics analyses and a systematic literature analysis of the (putative) function of the proteins encoded by the 85 top-ranked genes emerging from the five GWASs for ADHD, we describe a signaling network involved in neurite outgrowth that includes 45 of these proteins. The other 40 proteins could not be convincingly linked to the network. Corroborating evidence for the involvement of this gene network from chromosomal aberrations in humans, from animal models, and from psychopharmacological studies is also presented.
Method
Identification of ADHD Candidate Genes From GWASs
To date, the results of five GWASs of ADHD have been reported. Neale et al. (27) reported GWAS results for a categorical ADHD phenotype in 909 Caucasian case-parent trios that were collected as part of the International Multicenter ADHD Genetics study in children. Lasky-Su et al. (28) performed a GWAS on the same sample using quantitative phenotypes of ADHD.
Lesch et al. (29) conducted a GWAS for a categorically defined ADHD phenotype using pooled DNA from 343 ADHD-affected adults and 304 comparison subjects. Recently, Mick et al. (30) also reported the results of a GWAS of a categorical ADHD phenotype in a sample of 735 trios from three different U.S. clinical sites (30). In addition, the International Multicenter ADHD Genetics II consortium reported the results of a GWAS using a case-control design in 896 unrelated participants with ADHD and 2,455 comparison subjects (31). From these five GWASs, we compiled a list of top SNPs, taking into account SNPs located within exonic, intronic, or untranslated regions of genes and in association with the ADHD phenotype at p<1.00E-04 (Table 1).
| GWAS | SNP | p | Locus | Gene | Position Within Gene | Gene Size |
|---|---|---|---|---|---|---|
| Neale et al. (27)b | ||||||
| rs9676447 | 1.00E-05 | 19q13.33 | NUCB1 | intronic | 23 kb | |
| rs876477 | 2.70E-05 | 4p15.31 | KCNIP4 | intronic | 1,220 kb | |
| rs7657608 | 3.00E-05 | 4q32.3 | SPOCK3 | intronic | 501 kb | |
| rs3782309 | 5.50E-05 | 12p11.23 | ITPR2 | intronic | 498 kb | |
| rs922781 | 5.50E-05 | 15q22.2 | RORA | intronic | 741 kb | |
| rs12505502 | 5.60E-05 | 4q22.1 | FAM190A | intronic | 1,360 kb | |
| rs1062793 | 7.00E-05 | 6q14.1 | SH3BGRL2 | 3′ untranslated region | 72 kb | |
| rs703965 | 9.20E-05 | 10q22.3 | ZMIZ1 | intronic | 248 kb | |
| rs4149601 | 9.30E-05 | 18q21.31 | NEDD4L | intronic | 357 kb | |
| Lasky-Su et al. (28)c | ||||||
| rs6565113 | d | 16q23.3 | CDH13 | intronic | 948 kb | |
| rs552655 | d | 6p24.1 | GFOD1 | intronic | 130 kb | |
| rs4128767 | 1.28E-06 | 15q25.1 | IL16 | intronic | 117 kb | |
| rs11719664 | 2.48E-06 | 3p24.3 | ZNF385D | intronic | 955 kb | |
| rs7577925 | 2.55E-06 | 2q21.2 | NAP5 | intronic | 897 kb | |
| rs7495052 | 2.83E-06 | 15q26.1 | SLCO3A1 | intronic | 319 kb | |
| rs17281813 | 3.46E-06 | 16q12.1 | ZNF423 | intronic | 336 kb | |
| rs363512 | 3.89E-06 | 21q21.3 | GRIK1 | intronic | 403 kb | |
| rs8041675 | 3.98E-06 | 15q14 | MEIS2 | intronic | 210 kb | |
| rs17641078 | 4.73E-06 | 9p24.3 | DMRT2 | nonsynonymous, coding | 7 kb | |
| rs930421 | 5.64E-06 | 2p21 | MTA3 | 3′ untranslated region | 262 kb | |
| rs17651978 | 6.05E-06 | 3p13 | FOXP1 | intronic | 629 kb | |
| rs478597 | 8.08E-06 | 12q24.22 | NOS1 | intronic | 163 kb | |
| rs6791644 | 8.32E-06 | 3p14.2 | FHIT | intronic | 1,502 kb | |
| rs260461 | 8.38E-06 | 19q13.43 | ZNF544 | intronic | 35 kb | |
| rs2290416 | 8.51E-06 | 8q24.3 | NAPRT1 | synonymous, coding | 4 kb | |
| rs1202199 | 8.52E-06 | 6p22.3 | MBOAT1 | intronic | 110 kb | |
| Lesch et al. (29)e | ||||||
| rs864643 | 1.30E-08 | 3p22.2 | MOBP | 3′ untranslated region | 59 kb | |
| rs7995215 | 1.35E-08 | 13q31.3 | GPC6 | intronic | 1,180 kb | |
| rs11243897 | 5.63E-08 | 9q34.13 | C9orf98 | intronic | 153 kb | |
| rs7164335 | 1.30E-07 | 15q23 | ITGA11 | intronic | 130 kb | |
| rs220470 | 1.34E-07 | 17p13.2 | ITGAE | intronic | 87 kb | |
| rs10983238 | 1.37E-07 | 9q33.1 | ASTN2 | intronic | 990 kb | |
| rs2587695 | 2.73E-07 | 2q14.2 | PCDP1 | intronic | 118 kb | |
| rs2281597 | 5.41E-07 | 1p35.1 | CSMD2 | intronic | 652 kb | |
| rs10514604 | 8.06E-07 | 16q24.1 | ATP2C2 | intronic | 96 kb | |
| rs2677744 | 9.69E-07 | 15q26.1 | MAN2A2 | intronic | 18 kb | |
| rs2502731 | 1.58E-06 | 9q34.11 | DNM1 | intronic | 52 kb | |
| rs2199161 | 1.62E-06 | 5q13.2 | MAP1B | intronic | 100 kb | |
| rs10786284 | 1.96E-06 | 10q24.1 | TLL2 | intronic | 149 kb | |
| rs1555322 | 3.60E-06 | 20q11.22 | MMP24 | intronic | 50 kb | |
| rs16928529 | 3.90E-06 | 10q22.1 | UNC5B | intronic | 90 kb | |
| rs515910 | 4.36E-06 | 10q25.1 | C10orf79 | intronic | 102 kb | |
| rs2237349 | 4.63E-06 | 7p15.1 | CREB5 | intronic | 527 kb | |
| rs4964805 | 4.74E-06 | 12q23.3 | NT5DC3 | intronic | 71 kb | |
| rs3799977 | 4.90E-06 | 6p21.1 | SUPT3H | intronic | 569 kb | |
| rs11646411 | 7.40E-06 | 16q23.3 | CDH13 | intronic | 948 kb | |
| rs469727 | 7.54E-06 | 5q22.2 | REEP5 | intronic | 46 kb | |
| rs2241685 | 8.11E-06 | 2p25.3 | MYT1L | intronic | 542 kb | |
| rs2242073 | 8.27E-06 | 2q33.3 | CRYGC | intronic | 2 kb | |
| rs13395022 | 9.68E-06 | 2p12 | CTNNA2 | intronic | 1,460 kb | |
| rs11594082 | 1.00E-05 | 10q22.1 | CDH23 | intronic | 419 kb | |
| rs3893215 | 2.56E-05 | 11p15.1 | KCNC1 | intronic | 47 kb | |
| Mick et al. (30)f | ||||||
| rs2823819 | 6.70E-07 | 21q21.1 | C21orf34 | intronic | 539 kb | |
| rs11074889 | 6.71E-07 | 16p13.13 | EMP2 | intronic | 52 kb | |
| rs8074751 | 1.18E-06 | 17q24.1 | CCDC46 | intronic | 556 kb | |
| rs1859156 | 2.05E-06 | 4q22.3 | BMPR1B | intronic | 400 kb | |
| rs438259 | 3.71E-06 | 6p24.3 | AL024506.1 | intronic | 122 kb | |
| rs2602381 | 3.85E-06 | 2q37.1 | UGT1A9 | intronic | 101 kb | |
| rs10011926 | 8.07E-06 | 4q25 | ELOVL6 | intronic | 153 kb | |
| rs11953346 | 1.59E-05 | 5q15 | MCTP1 | intronic | 578 kb | |
| rs7702178 | 1.84E-05 | 5q35.1 | DUSP1 | intronic | 3 kb | |
| rs17023218 | 1.88E-05 | 3p24.1 | RBMS3 | intronic | 1,166 kb | |
| rs6561686 | 2.08E-05 | 13q14.3 | LECT1 | intronic | 37 kb | |
| International Multicenter ADHD Genetics II consortium (31)g | ||||||
| rs10823973 | 6.30E-07 | 10q21.1 | PRKG1 | intronic | 1,307 kb | |
| rs2291567 | 2.00E-06 | 7q32.1 | FLNC | intronic | 29 kb | |
| rs13238831 | 5.60E-06 | 7q32.1 | KCP | intronic | 48 kb | |
| rs11593241 | 5.70E-06 | 10q26.3 | TCERG1L | intronic | 219 kb | |
| rs12317552 | 9.10E-06 | 12q14.1 | PPM1H | 3′ untranslated region | 291 kb | |
| rs17151821 | 1.10E-05 | 7p21.3 | NXPH1 | intronic | 319 kb | |
| rs17095690 | 1.60E-05 | 1p31.1 | C1orf173 | intronic | 106 kb | |
| rs8045006 | 2.30E-05 | 16q23.3 | CDH13 | intronic | 948 kb | |
| rs1023556 | 2.60E-05 | 7p14.3 | AC005493.1 | intronic | 14 kb | |
| rs2394538 | 2.80E-05 | 10q22.1 | HK1 | intronic | 132 kb | |
| rs1278776 | 2.90E-05 | 13q34 | ATP11A | intronic | 197 kb | |
| rs17495366 | 3.10E-05 | 2p16.3 | NRXN1 | intronic | 1,114 kb | |
| rs906219 | 3.70E-05 | 10q22.1 | HKDC1 | nonsynonymous, coding | 47 kb | |
| rs1472147 | 4.10E-05 | 7q32.1 | ATP6V1F | intronic | 3 kb | |
| rs589943 | 4.90E-05 | 11q22.3 | DYNC2H1 | intronic | 370 kb | |
| rs2962393 | 5.60E-05 | 5q34 | GABRB2 | intronic | 260 kb | |
| rs846971 | 5.60E-05 | 6q21 | SOBP | intronic | 170 kb | |
| rs10445861 | 6.50E-05 | 2q14.3 | CNTNAP5 | intronic | 890 kb | |
| rs4261672 | 7.20E-05 | 2q22.2 | LRP1B | intronic | 1,900 kb | |
| rs6986578 | 7.30E-05 | 8q13.2 | PREX2 | intronic | 280 kb | |
| rs12090000 | 7.40E-05 | 1q25.2 | FAM5B | intronic | 111 kb | |
| rs1565630 | 8.00E-05 | 7q32.1 | CCDC136 | intronic | 31 kb | |
| rs4533267 | 8.30E-05 | 15q26.3 | ADAMTS17 | intronic | 370 kb | |
| rs2979307 | 9.00E-05 | 3q21.2 | OSBPL11 | intronic | 66 kb |
TABLE 1. The Top Single Nucleotide Polymorphisms (SNPs) From the Five Reported Genome-Wide Association Studies of Attention Deficit Hyperactivity Disorder (ADHD)a
Ingenuity and BiNGO Pathway Analyses
In order to detect significantly enriched gene categories in the selected ADHD candidate genes from the five GWASs, we performed analyses using the Ingenuity Pathway Analysis software package (http://www.ingenuity.com) and the BiNGO (Biological Network Gene Ontology) bioinformatics tool (33). We performed similar analyses on a set of top genes from four GWASs for diabetes type I and Crohn's disease and compared the results with those for ADHD in order to identify possible overrepresentation bias of large (brain-expressed) genes in the top findings of the ADHD GWASs.
The Ingenuity software package (http://www.ingenuity.com) uses the Ingenuity Knowledge Base, which is based on information from the published literature as well as many other sources, including gene expression and gene annotation databases, to assign genes to different groups and categories of functionally related genes. Each of these categories can be further divided into many subcategories. Ingenuity calculates single p values for the enrichment of each gene category using the right-tailed Fisher's exact test and taking into consideration both the total number of molecules from the analyzed data set and the total number of molecules that is linked to the same gene category according to the Ingenuity Knowledge Base. Furthermore, for each gene category, a multiple testing corrected p value of enrichment, calculated using the Benjamini-Hochberg correction, is provided.
The BiNGO tool also assigns genes to different functional categories, but these are specifically linked to gene ontology terms that group functionally related genes (33) and can be found in the Gene Ontology database ( http://www.geneontology.org/). The gene ontology terms can be further assigned to three main gene ontology subgroups or domains (i.e., cellular component, biological process, and molecular function). As with Ingenuity, the BiNGO tool provides single p values, calculated with the hypergeometric test and taking into consideration both the total number of molecules from the analyzed data set and the total number of molecules that is linked to the same gene ontology term, as well as multiple testing corrected p values, calculated using the Benjamini-Hochberg correction, for the enrichment of each gene ontology term (33).
In the presentation of the results of the Ingenuity and BiNGO analyses for ADHD, only categories/processes with significant enrichment (i.e., multiple testing corrected p<0.05) and containing more than one gene were taken into account. Only in the comparison between ADHD, diabetes type I, and Crohn's disease were the functional categories that only contain one gene also considered.
Literature Analysis
Subsequent to the bioinformatics analysis, we systematically searched the literature for the (proposed) function of all 85 proteins derived from the ADHD candidate genes using PubMed (http://www.ncbi.nlm.nih.gov/sites/entrez) and the UniProt Protein Knowledge Base (UniProtKB), a comprehensive catalog of protein sequences and functional annotations that is updated every 3 weeks and can be accessed online (http://www.uniprot.org/uniprot [34]). For each ADHD candidate, we first looked at the available information in UniProtKB, which in several cases already provided a general indication of the (putative) function of the gene/protein in question. Subsequently, we searched PubMed using the search terms “brain,” “neuron,” and “neurite” in combination with the name of each of the candidate genes/proteins from the GWASs. Lastly, guided by the literature we found, we searched PubMed for functional interactions between several of the candidate genes/proteins.
Results
Table 1 shows the 85 genes from the five GWASs for ADHD fulfilling the inclusion criteria of the present study. When more than one SNP in a single gene was found among the top findings, only the SNP yielding the lowest p value for association with (a quantitative phenotype of) ADHD is presented. As can be derived from Table 1, the only overlap between the top findings of the five published GWASs was for CDH13, in which (three different) SNPs were associated with ADHD at p<1.00E-04 in three GWASs (28, 29, 31).
Analysis of these 85 top ADHD candidate genes using the Ingenuity pathway software revealed a significant enrichment (p=4.11E-08 after correction for multiple testing) of the functional gene category neurological disease, with 44 of the 85 genes falling into this category (Table 2). Furthermore, analysis with the BiNGO bioinformatics tool revealed that the gene ontology processes (calcium) ion binding and hexokinase activity were significantly enriched in the 85 ADHD genes (2.94E-03<p<5.43E-03 and p=2.69E-02, respectively) (Table 2).
| Category | Genes | Significance | Adjusted Significance |
|---|---|---|---|
| Ingenuity Pathway Toolb | |||
| Neurological disease (44/85 genes) | ADAMTS17, ASTN2, ATP2C2, BMPR1B, C10ORF79, C21ORF34, C9ORF98, CCDC46,CDH13, CDH23, CNTNAP5,CREB5, CSMD2, CTNNA2, DYNC2H1, FAM190A, FHIT,FOXP1, GABRB2, GFOD1, GRIK1,ITGA11, ITPR2,KCNIP4, LRP1B, MAP1B, MCTP1, MEIS2, MOBP, MYT1L, NAPRT1,NOS1, NRXN1, NXPH1, PPM1H, PRKG1, RBMS3,RORA, SLCO3A1, SPOCK3, TLL2, UNC5B,ZNF385D,ZNF423 | 4.67E-10 | 4.11E-08 |
| Biological Network Gene Ontology toolc | |||
| Gene ontology termd | |||
| Ion binding | ADAMTS17, ATP11A,ATP2C2,BMPR1B,CDH13, CDH23,CREB5,DMRT2, DUSP1, FHIT, FOXP1, GABRB2,HK1,ITGA11,ITGAE, ITPR2, KCNC1,KCNIP4,LRP1B,MAN2A2, MCTP1,MMP24, MTA3,MYT1L,NOS1, NRXN1,NT5DC3, NUCB1,RORA,SOBP,SPOCK3, TLL2,ZMIZ1,ZNF385D,ZNF423,ZNF544 | 4.10E-06 | 2.94E-03 |
| Metal ion binding | ADAMTS17, ATP11A,ATP2C2,BMPR1B,CDH13, CDH23,CREB5,DMRT2, DUSP1, FHIT, FOXP1,HK1,ITGA11,ITGAE, ITPR2, KCNC1,KCNIP4, LRP1B,MAN2A2, MCTP1,MMP24, MTA3,MYT1L,NOS1, NRXN1,NT5DC3, NUCB1, RORA, SOBP,SPOCK3, TLL2,ZMIZ1,ZNF385D,ZNF423,ZNF544 | 7.45E-06 | 2.94E-03 |
| Cation binding | ADAMTS17,ATP2C2,BMPR1B,CDH13, CDH23,CREB5,DMRT2, DUSP1, FHIT, FOXP1,HK1,ITGA11,ITGAE, ITPR2, KCNC1,KCNIP4,LRP1B,MAN2A2, MCTP1,MMP24, MTA3,MYT1L,NOS1, NRXN1, NUCB1, RORA,SOBP,SPOCK3, TLL2,ZMIZ1,ZNF385D,ZNF423,ZNF544 | 1.19E-05 | 3.14E-03 |
| Calcium ion binding | ATP2C2,CDH13, CDH23,ITGA11,ITGAE, ITPR2,KCNIP4,LRP1B,MCTP1,MMP24,NRXN1, NUCB1, SPOCK3, TLL2 | 2.76E-05 | 5.43E-03 |
| Hexokinase activity | HK1, HKDC1 | 1.71E-04 | 2.69E-02 |
TABLE 2. Enrichment Analyses of the Top 85 Attention Deficit Hyperactivity Disorder (ADHD) Candidate Genes in the Five Reported Genome-Wide Association Studies of ADHDa
Further literature analysis of the (putative) function of the proteins encoded by the genes in the enriched BiNGO terms revealed that 21 out of 36 genes in the (calcium) ion binding category and two out of two hexokinase activity genes play a role in neurite outgrowth. Subsequently, we investigated the function of the entire set of 85 genes further and found that a total of 45 of the 85 included genes (indicated in bold in tables, figure legend, and throughout this article) fit into a neurodevelopmental network that is involved in directed neurite outgrowth, which is illustrated in Figure 1. The proposed signaling network links axonal guidance and neuronal cell adhesion proteins at the neuronal cell membrane with downstream acting adaptor proteins and neuronal cytoskeleton/extracellular matrix-associated proteins.
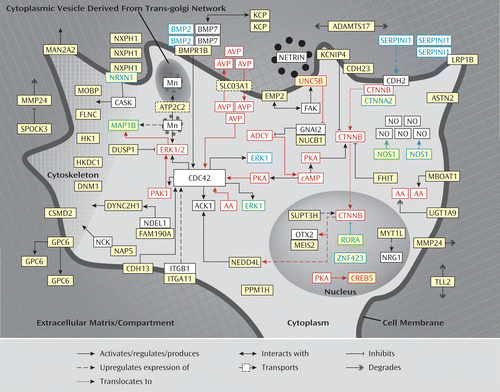
FIGURE 1. Schematic Representation of the ADHD Neurodevelopmental Signaling Network for Directed Neurite Outgrowtha
a The genes/proteins that emerged from the five reported genome-wide association studies for ADHD are indicated in yellow. The proteins encoded by genes found in copy number variations in patients with ADHD are indicated by a blue border. The proteins encoded by genes implicated in the etiology of ADHD through gene knockout studies in mice are indicated by a green border. The genes/proteins of which the expression and/or function is regulated by stimulants are indicated by a red border. Evidence placing the genes/proteins into this network is presented in the data supplement.
A detailed description of the evidence linking the genes in the network to neurite outgrowth is provided in the data supplement accompanying the online version of this article.
Involvement of the identified network in ADHD is also supported by the finding that several of the identified genes (i.e., CTNNA2, NRXN1, NAP5, SERPINI1, NOS1, ERK1, ZNF423, NEDD4L, and BMP2) are located within copy number variations-and are hence deleted or duplicated-in people with ADHD (20–22, 35–41) (Table 3, Figure 1).
Moreover, partial knockout models of the NOS1, ERK1, MAP1B, and RORA genes in mice provide further evidence of the involvement of these genes in ADHD etiology (Figure 1).
| Locus | Gene | Copy Number Variant | Phenotype | Additional Comments | Study |
|---|---|---|---|---|---|
| 2p12 | CTNNA2a | deleted | ADHD | Genome-wide copy number variant analysis | Elia et al. (20) |
| 2p16.3 | NRXN1a | deleted | ADHD | Genome-wide copy number variant analysis | Bradley et al. (35) |
| 2q21.2 | NAP5a | deleted | Attention deficit disorder | Baker et al. (36) | |
| 3q26.1 | SERPINI1 | duplicated | ADHD | Genome-wide copy number variant analysis | Elia et al. (20) |
| 3q26.1 | SERPINI1 | deleted | ADHD | Genome-wide copy number variant analysis | Lesch et al. (21) |
| 12q24.22 | NOS1a | duplicated | ADHD | Cappellacci et al. (37) | |
| 16p11.2 | ERK1 | duplicated | ADHD | Six/eight unrelated people with ADHD | Shinawi et al. (38) |
| 16p11.2 | ERK1 | duplicated | ADHD | Genome-wide copy number variant analysis | Williams et al. (22) |
| 16p11.2 | ERK1 | deleted | ADHD | Four/15 unrelated people with ADHD | Shinawi et al. (38) |
| 16p11.2 | ERK1 | deleted | ADHD | Genome-wide copy number variant analysis | Williams et al. (22) |
| 16q12.1 | ZNF423a | duplicated | ADHD | van den Berg et al. (39) | |
| 18q21.31 | NEDD4La | duplicated | Attention deficit disorder | Two siblings with attention deficit disorder | Ceccarini et al. (40) |
| 20p12.3 | BMP2 | deleted | Attention deficit disorder | Lalani et al. (41) |
TABLE 3. Genes Encoding Proteins From the Putative Attention Deficit Hyperactivity Disorder (ADHD) Network and Reported to be Deleted and/or Duplicated in (Genome-Wide) Copy Number Variations Among Individuals With ADHD
Mice in which the Nos1 and Erk1 genes have been knocked out display hyperactive behavior (42, 43), while, in contrast, Map1b- (44) and Rora-knockout mice (45) show hypoactivity.
Lastly, several proteins from the network appear to be under control of the stimulants methylphenidate and amphetamine that are used to treat ADHD symptoms (4, 5). Both stimulants have been shown to stimulate neurite outgrowth (46, 47) and directly or indirectly regulate the expression and/or function of several genes/proteins implicated in neurite outgrowth, including a number of genes/proteins from the identified network (Figure 1).
Methylphenidate exposure upregulates the expression of AVP (48) and CREB5 (49) (Figure 1). In addition, methylphenidate upregulates the expression of a large number of other genes not directly found in the identified network but functionally implicated in neurite outgrowth (50). Examples of these genes are MMP14, TIMP2, and TIMP3. The MMP14 gene encodes a neuronal extracellular matrix metalloproteinase, such as MMP24, and TIMP2 and TIMP3 encode two direct inhibitors of MMP14 (34, 50). Amphetamine upregulates the expression of UNC5B (51) and PAK1 (52) and downregulates the expression of CTNNB (53). Amphetamine also activates neuronal ERK1 and ERK2 (54) and increases the level of arachidonic acid in neocortical, extrapyramidal, and limbic brain regions (55). Deficiencies or imbalances of arachidonic acid have been shown to be associated with ADHD (56). Lastly, the adenylate cyclase/cAMP/protein kinase A/CREB-signaling cascade (including protein kinase A activating CREB5) is also activated by amphetamine (57).
An important potential bias of which to be aware in the analysis of the top findings from the reported GWASs of ADHD is the fact that large genes, which are often brain expressed, may be more likely found as associated with a phenotype in a GWAS as a result of chance, since more SNPs are present in these large genes (58). Indeed, the genes found among the top findings of the ADHD GWASs were considerably larger than the average gene size of the human genome ([Table 1] average size of 85 ADHD genes: 373 kb versus average gene size in human genome: 27 kb [59]).
If large genes are more likely found to be associated with a phenotype of a GWAS because of chance, this would be expected to be the case for GWASs of both nonpsychiatric disorders and ADHD, and therefore we compared the results of our analyses with the Ingenuity and BiNGO software tool results for the 20 top ADHD candidate genes with those for the top 20 candidate genes from four published GWASs for diabetes mellitus type I and Crohn's disease (Table 4, Table 5 [also see the data supplement]). This comparison showed that the neurological disease category was not enriched in Crohn's disease and only contained a single gene in diabetes type I (Table 4). In addition, the gene ontology processes that were enriched in the top GWAS findings were clearly different for the three diseases (Table 5).
| Category | Genes | Significanceb | Adjusted Significancec |
|---|---|---|---|
| ADHD | |||
| Neurological disease (13/20 genes) | ASTN2, ATP2C2, C21ORF34, C9ORF98, CCDC46, CDH13, CSMD2, GFOD1, ITGA11, MAP1B, MOBP, PRKG1, TLL2 | 1.53E-04 | 7.55E-03 |
| Biogenesis of lamellipodiad | CDH13 | 5.49E-03 | 2.10E-02 |
| Diabetes type I | |||
| Neurological disease (1/20 genes) | TH | 1.42E-02 | 3.02E-02 |
| Long-term depression | INS | 1.30E-03 | 8.11E-03 |
| Crohn's disease | |||
| Neurological disease (0/20 genes) |
TABLE 4. Nervous System-Related Gene Functional Categories Significantly Enriched in the Top 20 Candidate Genes From Genome-Wide Association Studies of Attention Deficit Hyperactivity Disorder (ADHD), Diabetes Type I, and Crohn's Diseasea
| Category | Genes | Significanceb | Adjusted Significancec |
|---|---|---|---|
| ADHDd | |||
| Gene ontology term | |||
| Regulation of chemotaxis | CDH13, IL16 | 3.58E-05 | 4.40E-03 |
| Regulation of behavior | CDH13, IL16 | 5.25E-05 | 4.40E-03 |
| Regulation of response to external stimulus | CDH13, IL16 | 2.77E-04 | 1.32E-02 |
| Integrin complex | ITGA11, ITGAE | 2.98E-04 | 1.32E-02 |
| Cell migration | CDH13, IL16, ITGA11 | 6.38E-04 | 2.34E-02 |
| Diabetes type Ie | |||
| Gene ontology term | |||
| Growth factor binding | ERBB3, IL7R, INS | 4.46E-05 | 2.27E-02 |
| Regulation of secretion | ERBB3, INS | 3.35E-04 | 3.54E-02 |
| Anion transport | C1QTNF6, CFTR, COL1A2 | 4.75E-04 | 3.54E-02 |
| Regulation of insulin receptor signaling | INS | 9.76E-04 | 3.54E-02 |
| Tyrosine 3-monooxygenase activity | TH | 9.76E-04 | 3.54E-02 |
| Crohn's diseasef | |||
| Gene ontology term | |||
| Protein kinase cascade | GCKR, LRRK2, MST1, STAT3, TNFSF15 | 5.10E-06 | 2.28E-03 |
| Intracellular signaling cascade | GCKR, LRRK2, MST1, NOD2, STAT3, TNFSF15 | 6.10E-04 | 2.42E-02 |
| Cell communication | ATG16L1, BSN, GCKR, ITLN1, LRRK2, MST1, NOD2, PTPN22, STAT3, TNFSF15 | 7.95E-04 | 2.42E-02 |
| Protein amino acid autophosphorylation | LRRK2, MST1 | 9.08E-04 | 2.42E-02 |
| Macrophage activation during immune response | SBNO2 | 9.76E-04 | 2.42E-02 |
TABLE 5. The Top Five Gene Functional Categories Significantly Enriched in the Top 20 Candidate Genes From Genome-Wide Association Studies of Attention Deficit Hyperactivity Disorder (ADHD), Diabetes Type I, and Crohn's Diseasea
Discussion
In the present study, we used both bioinformatics tools and manual literature mining to attempt to integrate the top findings of the currently available GWAS data for ADHD. We found that 45 of the 85 top ADHD candidate genes (or approximately 53%) fit into a network that is involved in neurite outgrowth. It should be noted that our decision to only include SNPs within exonic or intronic regions of genes and those with association at p<1.00E-04 is essentially arbitrary. By only including genes with SNPs in exons and introns, we have ignored the possible role of upstream and downstream regulatory sequences (61), which could have increased the number of (potentially relevant) genes to be included in the study. However, we do not expect that this would have altered the general tendency in our results (i.e., the considerable overrepresentation of neurite outgrowth-related genes).
Surprisingly, the enrichment of neurite outgrowth-related genes was not reflected by the finding of gene functional categories and/or gene ontology terms directly related to neurite outgrowth with the bioinformatics tools used in this study (Table 2). This shows the limitations of the currently available bioinformatics software, which is a result of the current incompleteness of the annotation of the human genome.
Therefore, use of bioinformatics tools should always be accompanied by manual and systematic literature and database mining in order to fully understand the processes involved in the etiology of multifactorial disorders. Moreover, the enrichment of neurite outgrowth-related genes in the ADHD GWAS data (53%) should be viewed from the perspective that only 3% (N=576) of all currently annotated human genes (N=18,589) are involved in neurogenesis (gene ontology term: GO:0022008).
Several additional lines of evidence suggest that the identified network provides an important contribution to the etiology of ADHD. Apart from implicating the same genes as the GWASs, these other lines of evidence point to involvement of several other genes (and their corresponding proteins) that were not found (at the p<1.00E-04 level) in the GWASs but that functionally connect the genes from the GWAS findings.
First, nine proteins that act in the identified neurite outgrowth network are encoded by genes that were also found in copy number variations in people with ADHD (20–22, 35–41) (Table 3). Second, knockout mouse models of four network genes reveal ADHD-related behavior (42–45). Third, and most importantly, the expression and/or function of a number of genes/proteins in the identified network is regulated by the stimulants methylphenidate (48–50) and amphetamine (51–55, 57), which are the most commonly used psychopharmacological treatments for ADHD symptoms (4, 5) and directly regulate neurite outgrowth (46, 47). The latter finding might not only increase our understanding of the working mechanism of these drugs but also provide directions for future research into new and more effective psychopharmacological ADHD treatments.
The fact that stimulants regulate neurite outgrowth potentially implicates a number of classic ADHD candidate genes in our network. Methylphenidate and amphetamine mediate effects on both dopaminergic and serotonergic neurotransmission in the brain (4, 5, 62). In this respect, the genes encoding the SLC6A3, DRD4, DRD5, SLC6A4, and HTR1B proteins could be putatively linked to the identified network. In addition, the link of these classic candidate genes with the neurite outgrowth network may help to localize the network to dopaminergic and serotonergic brain regions.
The findings from this study fit very well with literature about disturbed brain structure and function in ADHD patients (7–10) and, even more so, with recent reports describing aberrant structural and functional brain connectivity in these patients (11–17). Because differences in brain connectivity have been shown to underlie interindividual variability in complex cognition-related processes, including ADHD (63, 64), relatively minor alterations in neurite outgrowth efficiency or direction may provide a major contribution to the cognitive deficits observed in ADHD.
A possibly important bias in our study may arise from the fact that the brain-expressed genes in the top findings of the ADHD GWASs are substantially larger than the average gene size in the human genome, and seven of the nine very large genes (i.e., genes with a size of >1 Mb) from the list of 85 ADHD genes indeed encode proteins that fit into the proposed ADHD network. It has been argued that the overrepresentation of large genes in the top findings of GWASs might result from bias because one is more likely to find an associated SNP in a larger, usually brain-expressed gene (58). If this is the case, it would be expected that similar results would be found in GWASs of polygenic disorders that are assumed not to primarily originate in the brain, such as diabetes mellitus and Crohn's disease. However, our Ingenuity and BiNGO analyses on the 20 top candidate genes from the GWASs for ADHD, diabetes mellitus type I, and Crohn's disease (Table 4, Table 5) show that this bias does not likely explain the findings for ADHD. In this respect, we would like to submit that the strong enrichment of neurite outgrowth-related genes in the top findings from the five ADHD GWASs, with 45 out of 85 genes fitting into our proposed network, also argues against the fact that these genes are spurious association findings. Nevertheless, we cannot completely exclude the possibility that some of the genes placed in the network still might have been chance findings.
Is the involvement of neurite outgrowth genes specific to the etiology of ADHD? This seems rather unlikely, since we recently also found neurite outgrowth to be implicated in dyslexia (65). Following an approach similar to the one used in the present study, we found that 10 of 14 dyslexia candidate genes reported to date (i.e., DCDC2, DIP2A, DOCK4, DYX1C1, FMR1, GTF2I, KIAA0319, KIAA0319L, ROBO1, S100B ) fit into a protein network that is involved in neurite outgrowth and neuronal migration. As shown in Figure 2 and explained in further detail in the data supplement, the proteins encoded by six of these genes (i.e., DCDC2, DYX1C1, FMR1, TFII-I [encoded by GTF2I], ROBO1, and S100B) fit directly into the identified network for ADHD. It is also interesting to note that decreased serum concentrations of S100B correlate with increasing hyperactivity symptoms in children with ADHD relative to normal comparison subjects (66) and that knockout mouse models of the FMR1 (67) and GTF2I (68) genes show ADHD-like behavior. One possible explanation for the observed genetic overlap between ADHD and dyslexia is that these neurodevelopmental disorders are highly comorbid and genetically correlated with one another (69–71). The same also holds true for autism (72), and recent evidence suggests involvement of neurite outgrowth genes in this neurodevelopmental disorder as well (73). The different functional consequences of disturbed or abnormal neurite outgrowth in the different brain regions that are most affected in ADHD, dyslexia, and autism, respectively, could help to explain why a disturbance of the same neurodevelopmental process could lead to these partially overlapping but still distinct clinical phenotypes.
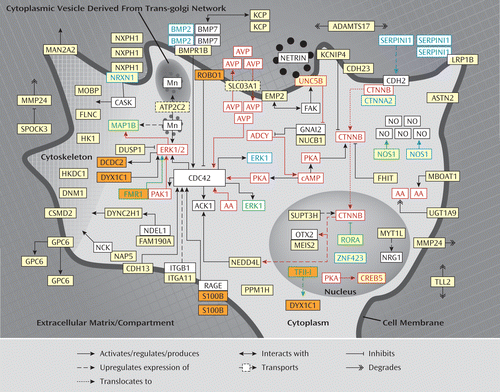
FIGURE 2. Schematic Representation of How Six Neurite Outgrowth-Related Dyslexia Candidate Genes Can be Directly Placed in the Signaling Network for ADHDa
a The six neurite outgrowth-related dyslexia candidate genes (65) that can be placed in the signaling network for ADHD are indicated in orange. The genes/proteins that emerged from the five reported genome-wide association studies for ADHD are indicated in yellow. The proteins encoded by genes found in copy number variations in patients with ADHD are indicated by a blue border. The proteins encoded by genes implicated in the etiology of ADHD through gene knockout studies in mice are indicated by a green border. The genes/proteins of which the expression and/or function is regulated by stimulants are indicated by a red border. Evidence placing the six dyslexia candidate genes/proteins into the ADHD network is presented in the data supplement.
Several genes that are major players in the identified neurite outgrowth network have not been directly observed in the top findings of the GWASs. An example of such a gene is ERK1, which not only has an important function in the network but has also been implicated in ADHD through its location in ADHD-related copy number variations (22, 38) as well as the fact that Erk1-knockout mice display hyperactive behavior (43) and that amphetamine directly activates neuronal ERK1 (54). Other examples are CTNNB and CDC42. These and other genes from the network may be strong candidates for future association studies.
Lastly, our data make a compelling case for the use of polygenic types of association analyses to explain a higher percentage of the heritability of ADHD through the GWAS findings. In this respect, all the individually associated genes from the identified neurite outgrowth network could be fitted simultaneously into one polygenic risk test, as illustrated in the recent study by Yang et al. (74) of all SNPs that have been individually associated in GWASs of variation in human height as well as in two other recent studies applying similar approaches to GWAS data for schizophrenia (75) and functional gene groups for cognitive ability in ADHD (76).
In conclusion, by integrating the top findings of the five GWASs for ADHD with copy number variation data, (psychopharmacologically induced) expression data, and data from animal studies, we have identified a protein signaling network for ADHD that results in directed neurite outgrowth. Systems biology approaches like those used in this study are needed to yield genetic findings that are useful for clinical purposes, such as the prediction of disease prognosis and the identification of new treatment targets and strategies.
1. : The worldwide prevalence of ADHD: a systematic review and metaregression analysis. Am J Psychiatry 2007; 164:942–948Link, Google Scholar
2. : The prevalence and correlates of adult ADHD in the United States: results from the National Comorbidity Survey Replication. Am J Psychiatry 2006; 163:716–723Link, Google Scholar
3. : Cross-national prevalence and correlates of adult attention-deficit hyperactivity disorder. Br J Psychiatry 2007; 190:402–409Crossref, Medline, Google Scholar
4. : The neurobiology of attention deficit/hyperactivity disorder. Eur J Paediatr Neurol 2009; 13:299–304Crossref, Medline, Google Scholar
5. : Mechanism of action of agents used in attention-deficit/hyperactivity disorder. J Clin Psychiatry 2006; 67 (suppl 8):32–38Crossref, Medline, Google Scholar
6. : Evaluating dopamine reward pathway in ADHD: clinical implications. JAMA 2009; 302:1084–1091Crossref, Medline, Google Scholar
7. : Developmental brain anomalies in children with attention-deficit hyperactivity disorder. J Child Neurol 2000; 15:102–108Crossref, Medline, Google Scholar
8. : Meta-analysis of structural imaging findings in attention-deficit/hyperactivity disorder. Biol Psychiatry 2007; 61:1361–1369Crossref, Medline, Google Scholar
9. : Abnormal cerebral cortex structure in children with ADHD. Hum Brain Mapp 2009; 30:175–184Crossref, Medline, Google Scholar
10. : Development of cortical asymmetry in typically developing children and its disruption in attention-deficit/hyperactivity disorder. Arch Gen Psychiatry 2009; 66:888–896Crossref, Medline, Google Scholar
11. : Attention-deficit/hyperactivity disorder: a preliminary diffusion tensor imaging study. Biol Psychiatry 2005; 57:448–455Crossref, Medline, Google Scholar
12. : Attention and executive systems abnormalities in adults with childhood ADHD: A DT-MRI study of connections. Cereb Cortex 2008; 18:1210–1220Crossref, Medline, Google Scholar
13. : Diffusion tensor imaging study of white matter fiber tracts in pediatric bipolar disorder and attention-deficit/hyperactivity disorder. Biol Psychiatry 2009; 65:586–593Crossref, Medline, Google Scholar
14. : Alerting deficits in children with attention deficit/hyperactivity disorder: event-related fMRI evidence. Brain Res 2008; 1219:159–168Crossref, Medline, Google Scholar
15. : Altered small-world brain functional networks in children with attention-deficit/hyperactivity disorder. Hum Brain Mapp 2009; 30:638–649Crossref, Medline, Google Scholar
16. : The restless brain: attention-deficit hyperactivity disorder, resting-state functional connectivity, and intrasubject variability. Can J Psychiatry 2009; 54:665–672Crossref, Medline, Google Scholar
17. : Functional disconnection of frontal cortex and visual cortex in attention-deficit/hyperactivity disorder. Biol Psychiatry 2010; 67:617–623Crossref, Medline, Google Scholar
18. : Attention-deficit hyperactivity disorder. Lancet 2005; 366:237–248Crossref, Medline, Google Scholar
19. : Molecular genetics of attention-deficit/hyperactivity disorder. Biol Psychiatry 2005; 57:1313–1323Crossref, Medline, Google Scholar
20. : Rare structural variants found in attention-deficit hyperactivity disorder are preferentially associated with neurodevelopmental genes. Mol Psychiatry 2010; 15:637–646Crossref, Medline, Google Scholar
21. : Genome-wide copy number variation analysis in attention-deficit/hyperactivity disorder: association with neuropeptide Y gene dosage in an extended pedigree. Mol Psychiatry (Epub ahead of print, March 23, 2010)Google Scholar
22. : Rare chromosomal deletions and duplications in attention-deficit hyperactivity disorder: a genome-wide analysis. Lancet 2010; 376:1401–1408Crossref, Medline, Google Scholar
23. : Meta-analysis of genome-wide linkage scans of attention deficit hyperactivity disorder. Am J Med Genet B Neuropsychiatr Genet 2008; 147B:1392–1398Crossref, Medline, Google Scholar
24. : Suggestive linkage of ADHD to chromosome 18q22 in a young genetically isolated Dutch population. Eur J Hum Genet 2009; 17:958–966Crossref, Medline, Google Scholar
25. : Candidate gene studies of ADHD: a meta-analytic review. Hum Genet 2009; 126:51–90Crossref, Medline, Google Scholar
26. : Meta-analysis shows significant association between dopamine system genes and attention deficit hyperactivity disorder (ADHD). Hum Mol Genet 2006; 15:2276–2284Crossref, Medline, Google Scholar
27. : Genome-wide association scan of attention deficit hyperactivity disorder. Am J Med Genet B Neuropsychiatr Genet 2008; 147B:1337–1344Crossref, Medline, Google Scholar
28. : Genome-wide association scan of quantitative traits for attention deficit hyperactivity disorder identifies novel associations and confirms candidate gene associations. Am J Med Genet B Neuropsychiatr Genet 2008; 147B:1345–1354Crossref, Medline, Google Scholar
29. : Molecular genetics of adult ADHD: converging evidence from genome-wide association and extended pedigree linkage studies. J Neural Transm 2008; 115:1573–1585Crossref, Medline, Google Scholar
30. : Family-based genome-wide association scan of attention-deficit/hyperactivity disorder. J Am Acad Child Adolesc Psychiatry 2010; 49:898–905Crossref, Medline, Google Scholar
31. ;
32. : Genome-wide association studies in ADHD. Hum Genet 2009; 126:13–50Crossref, Medline, Google Scholar
33. : BiNGO: a Cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 2005; 21:3448–3449Crossref, Medline, Google Scholar
34.
35. : Hotspots of large rare deletions in the human genome. PLoS One 2010; 5:e9401Crossref, Medline, Google Scholar
36. : Chromosome 2 interstitial deletion (del[2][q14.1q21]) associated with connective tissue laxity and an attention deficit disorder. J Med Genet 2001; 38:493–496Crossref, Medline, Google Scholar
37. : De novo pure 12q22q24.33 duplication: first report of a case with mental retardation, ADHD, and Dandy-Walker malformation. Am J Med Genet A 2006; 140:1203–1207Crossref, Medline, Google Scholar
38. : Recurrent reciprocal 16p11.2 rearrangements associated with global developmental delay, behavioral problems, dysmorphism, epilepsy, and abnormal head size. J Med Genet 2010; 47:332–341Crossref, Medline, Google Scholar
39. : Investigation of a patient with a partial trisomy 16q including the fat mass and obesity associated gene (FTO): fine mapping and FTO gene expression study. Am J Med Genet A 2010; 152A:630–637Crossref, Medline, Google Scholar
40. : Duplication 18q21.31-q22.2. Am J Med Genet A 2007; 143:343–348Crossref, Medline, Google Scholar
41. : 20p12.3 microdeletion predisposes to Wolff-Parkinson-White syndrome with variable neurocognitive deficits. J Med Genet 2009; 46:168–175Crossref, Medline, Google Scholar
42. : Abnormal social behavior, hyperactivity, impaired remote spatial memory, and increased D1-mediated dopaminergic signaling in neuronal nitric oxide synthase knockout mice. Mol Brain 2009; 2:19Crossref, Medline, Google Scholar
43. : The extracellular signal-regulated kinase pathway contributes to the control of behavioral excitement. Mol Psychiatry 2009; 14:448–461Crossref, Medline, Google Scholar
44. : Mice deficient in microtubule-associated protein MAP1B show a distinct behavioral phenotype and altered retina function. Behav Brain Res 2005; 164:188–196Crossref, Medline, Google Scholar
45. : Regional brain variations of cytochrome oxidase activity and motor co-ordination in staggerer mutant mice. Neuroscience 2000; 95:903–911Crossref, Medline, Google Scholar
46. : 3,4-Methylenedioxy-N-methamphetamine (ecstasy) promotes the survival of fetal dopamine neurons in culture. Neuropharmacology 2008; 55:851–859Crossref, Medline, Google Scholar
47. : Repeated amphetamine treatment induces neurite outgrowth and enhanced amphetamine-stimulated dopamine release in rat pheochromocytoma cells (PC12 cells) via a protein kinase C- and mitogen activated protein kinase-dependent mechanism. J Neurochem 2003; 87:1546–1557Crossref, Medline, Google Scholar
48. : Methylphenidate-induced motor activity in rats: modulation by melatonin and vasopressin. Pharmacol Biochem Behav 2003; 75:67–73Crossref, Medline, Google Scholar
49. : Altered responsiveness to cocaine in rats exposed to methylphenidate during development. Nat Neurosci 2002; 5:13–14Crossref, Medline, Google Scholar
50. : Short-term effects of adolescent methylphenidate exposure on brain striatal gene expression and sexual/endocrine parameters in male rats. Ann N Y Acad Sci 2006; 1074:52–73Crossref, Medline, Google Scholar
51. : Regulation of netrin-1 receptors by amphetamine in the adult brain. Neuroscience 2007; 150:764–773Crossref, Medline, Google Scholar
52. : Transient enhanced expression of CDK5 activator p25 after acute and chronic d-amphetamine administration. Ann N Y Acad Sci 2008; 1139:89–102Crossref, Medline, Google Scholar
53. : The effects of antipsychotics on beta-catenin, glycogen synthase kinase-3 and dishevelled in the ventral midbrain of rats. J Neurochem 2005; 95:513–525Crossref, Medline, Google Scholar
54. : CaMKII regulates amphetamine-induced ERK1/2 phosphorylation in striatal neurons. Neuroreport 2002; 13:1013–1016Crossref, Medline, Google Scholar
55. : D-amphetamine stimulates D2 dopamine receptor-mediated brain signaling involving arachidonic acid in unanesthetized rats. J Cereb Blood Flow Metab 2006; 26:1378–1388Crossref, Medline, Google Scholar
56. : Significance of long-chain polyunsaturated fatty acids (PUFAs) for the development and behaviour of children. Eur J Pediatr 2010; 169:149–164Crossref, Medline, Google Scholar
57. : Cocaine-amphetamine-regulated transcript expression in the rat nucleus accumbens is regulated by adenylyl cyclase and the cyclic adenosine 5′-monophosphate/protein kinase: a second messenger system. J Pharmacol Exp Ther 2006; 317:454–461Crossref, Medline, Google Scholar
58. : Gene ontology analysis of GWA study data sets provides insights into the biology of bipolar disorder. Am J Hum Genet 2009; 85:13–24Crossref, Medline, Google Scholar
59. : Human Molecular Genetics. New York, Garland Publishing, 2004, p 253Google Scholar
60. : CDC42 stimulates neurite outgrowth and formation of growth cone filopodia and lamellipodia. J Neurobiol 2000; 43:352–364Crossref, Medline, Google Scholar
61. : High-resolution mapping of expression-QTLs yields insight into human gene regulation. PLoS Genet 2008; 4:e1000214Crossref, Medline, Google Scholar
62. : Comparison of the monoamine transporters from human and mouse in their sensitivities to psychostimulant drugs. BMC Pharmacol 2006; 6:6Crossref, Medline, Google Scholar
63. : Genetics of brain fiber architecture and intellectual performance. J Neurosci 2009; 29:2212–2224Crossref, Medline, Google Scholar
64. : Sex differences in the functional neuroanatomy of working memory in adults with ADHD. Am J Psychiatry 2010; 167:86–94Link, Google Scholar
65. : A theoretical molecular network for dyslexia: integrating available genetic findings. Mol Psychiatry 2010 (Epub ahead of print October 19, 2010)Medline, Google Scholar
66. : Attention-deficit hyperactivity disorder (ADHD) and glial integrity: an exploration of associations of cytokines and kynurenine metabolites with symptoms and attention. Behav Brain Funct 2010; 6:32Crossref, Medline, Google Scholar
67. : Attentional dysfunction, impulsivity, and resistance to change in a mouse model of fragile X syndrome. Behav Neurosci 2006; 120:1367–1379Crossref, Medline, Google Scholar
68. : Essential role of the N-terminal region of TFII-I in viability and behavior. BMC Med Genet 2010; 1161Google Scholar
69. : Hyperactivity and spelling disability: testing for shared genetic aetiology. J Child Psychol Psychiatry 1993; 34:1137–1152Crossref, Medline, Google Scholar
70. : Reading disability and hyperactivity: evidence for a common genetic etiology. Dev Neuropsychol 1995; 11:323–335Crossref, Google Scholar
71. : Hyperactivity and reading disability: a longitudinal study of the nature of the association. J Child Psychol Psychiatry 1999; 40:1039–1050Crossref, Medline, Google Scholar
72. : Shared heritability of attention-deficit/hyperactivity disorder and autism spectrum disorder. Eur Child Adolesc Psychiatry 2010; 19:281–295Crossref, Medline, Google Scholar
73. : Systematic resequencing of X-chromosome synaptic genes in autism spectrum disorder and schizophrenia. Mol Psychiatry 2010 (Epub ahead of print, May 18, 2010)Google Scholar
74. : Common SNPs explain a large proportion of the heritability for human height. Nat Genet 2010; 42:565–569Crossref, Medline, Google Scholar
75. : Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 2009; 460:748–752Crossref, Medline, Google Scholar
76. : Functional gene group analysis reveals a role of synaptic heterotrimeric G proteins in cognitive ability. Am J Hum Genet 2010; 86:113–125Crossref, Medline, Google Scholar



