Interrogating the Genetic Determinants of Tourette’s Syndrome and Other Tic Disorders Through Genome-Wide Association Studies
Abstract
Objective:
Tourette’s syndrome is polygenic and highly heritable. Genome-wide association study (GWAS) approaches are useful for interrogating the genetic architecture and determinants of Tourette’s syndrome and other tic disorders. The authors conducted a GWAS meta-analysis and probed aggregated Tourette’s syndrome polygenic risk to test whether Tourette’s and related tic disorders have an underlying shared genetic etiology and whether Tourette’s polygenic risk scores correlate with worst-ever tic severity and may represent a potential predictor of disease severity.
Methods:
GWAS meta-analysis, gene-based association, and genetic enrichment analyses were conducted in 4,819 Tourette’s syndrome case subjects and 9,488 control subjects. Replication of top loci was conducted in an independent population-based sample (706 case subjects, 6,068 control subjects). Relationships between Tourette’s polygenic risk scores (PRSs), other tic disorders, ascertainment, and tic severity were examined.
Results:
GWAS and gene-based analyses identified one genome-wide significant locus within FLT3 on chromosome 13, rs2504235, although this association was not replicated in the population-based sample. Genetic variants spanning evolutionarily conserved regions significantly explained 92.4% of Tourette’s syndrome heritability. Tourette’s-associated genes were significantly preferentially expressed in dorsolateral prefrontal cortex. Tourette’s PRS significantly predicted both Tourette’s syndrome and tic spectrum disorders status in the population-based sample. Tourette’s PRS also significantly correlated with worst-ever tic severity and was higher in case subjects with a family history of tics than in simplex case subjects.
Conclusions:
Modulation of gene expression through noncoding variants, particularly within cortico-striatal circuits, is implicated as a fundamental mechanism in Tourette’s syndrome pathogenesis. At a genetic level, tic disorders represent a continuous spectrum of disease, supporting the unification of Tourette’s syndrome and other tic disorders in future diagnostic schemata. Tourette’s PRSs derived from sufficiently large samples may be useful in the future for predicting conversion of transient tics to chronic tic disorders, as well as tic persistence and lifetime tic severity.
Tourette’s syndrome is a complex neuropsychiatric disorder that occurs along a phenotypic spectrum that also includes chronic (persistent) motor or vocal tic disorder (chronic tics) and transient (provisional) tic disorder (1). Although Tourette’s syndrome is highly heritable (2), variants in known Tourette’s risk genes (e.g., CNTN6, NRXN1, SLITRK1, HDC, and CELSR3) account for less than 2% of affected individuals (3–6). Tourette’s syndrome is highly polygenic, with a demonstrated role for multiple common genetic variants of small effect distributed widely across the genome (7). Thus, genome-wide association studies (GWASs) (8) will be of benefit in further elucidating the underlying genetic etiology of the disorder.
To date, only one Tourette’s GWAS has been published (9). Although no single-nucleotide polymorphisms (SNPs) met criteria for genome-wide significance (p<5×10−8), in aggregate, the top SNPs (p values <1×10−3) were enriched for expression quantitative trait loci (eQTLs) in the frontal cortex and for methylation quantitative trait loci (mQTLs) in the cerebellum, indicating that a significant proportion of these variants have biological relevance to Tourette’s syndrome, and perhaps also to other tic disorders. However, as with other neuropsychiatric disorders, much larger sample sizes are needed to elucidate the disorder’s genetic underpinnings. Here, we report the results of a GWAS meta-analysis from the Psychiatric Genomics Consortium (PGC) Tourette Syndrome Workgroup in a sample that is nearly four times larger than the initial GWAS (9). We also probed aggregated Tourette’s syndrome polygenic risk to test two specific hypotheses: whether Tourette’s and related tic disorders have an underlying shared genetic etiology and whether Tourette’s polygenic risk scores correlate with worst-ever tic severity and may represent a future potential predictor of disease severity.
Methods
Study Subjects
The primary GWAS meta-analysis consisted of four European ancestry (EU) GWAS data sets: 1) 969 case subjects and 3,923 ancestry-matched control subjects from the initial Tourette’s syndrome GWAS (GWAS1) (9); 2) 2,711 additional EU Tourette’s case subjects (4) and 3,762 ancestry-matched control subjects (GWAS2); 3) Tourette’s probands from GWAS1 and one or more of their Tourette’s-affected family members (10) (N=548) plus 597 ancestry-matched control subjects (GWAS2 FAM); and 4) 591 independent EU Tourette’s probands from the Tourette International Collaborative Genetics (TIC) study (11) and 1,206 unselected ancestry-matched control subjects. Genotyping details are provided in Tables S1 and S2 in the online supplement.
GWAS1.
A total of 1,285 EU case subjects (number of cases from the GWAS1 paper) were collected from Tourette’s syndrome specialty clinics in the United States, Canada, the United Kingdom, and the Netherlands or through recruitment from the membership of the Tourette Association of America. Tourette’s diagnoses were based on DSM-IV-TR criteria plus observation of tics by an experienced clinician. After removing the subjects who were relatives or duplicates of subjects in GWAS2 or GWAS2 FAM, a total of 969 cases were retained for analysis. A total of 3,923 control subjects were identified primarily from previously genotyped unselected population control subjects and were ancestry-matched to the case subjects (9).
GWAS2.
A total of 2,871 EU case subjects with DSM-5 Tourette’s syndrome were identified by e-mail or online recruitment combined with validated, web-based phenotypic assessments (12, 13) (N=1,264) or from Tourette’s syndrome specialty clinics in the United States, Canada, and Europe (N=1,607) (see the Supplemental Methods section in the online supplement). All subjects were genotyped at the UCLA Neuroscience Genomics Core. After quality control, 2,711 case subjects were retained for analysis.
GWAS2 FAM.
The family sample consisted of 548 probands and first-degree relatives with Tourette’s syndrome from 207 independent families (10). A total of 175 probands came from the original Tourette’s GWAS1 sample; these case subjects were removed from the GWAS1 analysis along with ancestry-matched control subjects and reanalyzed with the family-based sample. Thirty-two Tourette’s probands and 341 additional Tourette’s-affected family members (total N=373) were genotyped along with the GWAS2 case-control sample. A total of 597 ancestry-matched control subjects were selected from a pool of previously genotyped control subjects (see the Supplemental Methods section in the online supplement).
TIC.
The TIC Genetics sample consisted of 591 probands, 579 of whom met DSM-5 criteria for Tourette’s syndrome and 12 of whom met criteria for DSM-5 chronic motor or vocal tic disorder (see Tables S1 and S2 in the online supplement).
Control subjects.
A total of 6,920 EU control subjects were obtained from cohorts of previously genotyped unselected population control subjects for the GWAS2, GWAS2 FAM, and TIC Genetics analyses; an additional 595 EU control subjects were genotyped with the Tourette’s case subjects at the UCLA Neuroscience Genomics Core (see Table S1 and the Supplemental Methods section in the online supplement).
deCODE.
An independent case-control replication sample from Iceland (deCODE genetics, Reykjavik) consisted of 706 Icelandic Tourette’s syndrome case subjects and 466 case subjects with other tic disorders (chronic tics or unspecified tic disorder) (see the Supplemental Methods section in the online supplement). A total of 127,164 unscreened population-matched control subjects were also available, of whom 6,068 were screened and reported no lifetime subclinical motor or vocal tics. Case and control subjects were genotyped at deCODE on Illumina SNP arrays (see the Supplemental Methods section).
Participants age 18 and older provided written informed consent; individuals under 18 gave assent, and parental permission was obtained. The study was approved by the human subjects committees at all participating sites.
Quality Control
Genotyping quality control was performed in PLINK, version 1.9 (14) (see the Supplemental Methods section). Duplicates and relatives were identified using genome-wide identity-by-descent estimates, and one member of each duplicate or relative pair was removed from the case-control sample. Relative pairs in which both individuals had a Tourette’s diagnosis were removed from the case-control sample and moved to the family-based analysis.
Population stratification was assessed through multidimensional scaling (MDS) analysis; individuals of non-European ancestry and extreme outliers on each of the MDS components were removed (see the Supplemental Methods section and Figure S1 in the online supplement). Case-control matching was verified across all MDS components. The final post–quality control GWAS2 sample contained 2,711 case subjects, 3,762 ancestry-matched control subjects, and 550,550 SNPs; the final GWAS2 FAM sample contained 548 case subjects and family members, 597 ancestry-matched control subjects, and 236,748 SNPs. The final TIC sample included 591 case subjects, 1,206 ancestry-matched control subjects, and 581,774 SNPs (see Table S2 in the online supplement).
Imputation and Genome-Wide Association
SNP imputation was conducted on all genotype data for the primary meta-analyses using the 1000 Genomes Project phase 1 integrated haplotypes (December 2013 release, with singleton sites removed) as the reference panel (15). SHAPEIT was used to phase genotype data, followed by imputation with IMPUTE, version 2. SNPs with INFO score <0.6 or certainty <0.9 were excluded.
Genome-wide association tests were performed on the imputed dosage data of the GWAS2 and TIC samples separately in PLINK 1.9, using logistic regression under an additive model with the first four MDS components and any additional MDS components associated with Tourette’s case-control status at p<0.05 included as covariates. A linear mixed model was used for the GWAS2 FAM association analysis in MMM, version 1.0 (16), to control for familial relatedness. GWAS1 samples were reimputed as described above; association tests were performed in four ancestry-based strata: nonisolate European (GWAS1_EU), Ashkenazi Jewish (GWAS1_AJ), French Canadian (GWAS1_FC), and GWAS1 TIC (GWAS1_TIC) (see Table S2).
A primary GWAS meta-analysis was conducted on the GWAS1, GWAS2, GWAS2 FAM, and TIC data sets using the inverse-variance method in METAL (17). Heterogeneity was assessed with Cochran’s I2 statistic. The genomic control factor (λ) was calculated for each individual GWAS and for the overall meta-analysis using all SNPs with minor allele frequency (MAF) >0.01 to identify residual population stratification or systematic technical artifact (see Figure S2 in the online supplement). GWAS summary statistics were subjected to linkage disequilibrium (LD) score regression (LDSC) analyses on high-quality common SNPs (INFO score >0.9 and MAF >0.01) to examine the LDSC intercept as a more specific measure of inflation of the GWAS test statistic (18) due to residual artifact or stratification. The genome-wide significance threshold for the GWAS (19, 20) was set at a p value of 5.0×10−8.
Heritability Estimation
Tourette’s syndrome SNP-based heritability was estimated on the liability scale, assuming a population prevalence of 0.8% (21), using both LDSC (18) and, in the GWAS1 and GWAS2 samples after excluding Ashkenazi Jewish and French Canadian samples, genotype-level data in a linear mixed model framework (7). To compare the relative polygenic burden of Tourette’s samples collected with different ascertainment methods, the Tourette’s GWAS1 and GWAS2 data sets were separated into three groups: GWAS1 case subjects (25% from affected sibling-pair families) (10); GWAS2 case subjects recruited through Tourette’s syndrome specialty clinics; and GWAS2 case subjects recruited via e-mail from the membership of the Tourette Association of America and assessed with a web-based phenotyping instrument (12). After additional stringent quality control of SNPs and samples (see the Supplemental Methods section), the SNP-based heritability of each ascertainment group was estimated both separately and jointly.
Partitioned heritability analyses were conducted using LDSC to evaluate enrichment of Tourette’s SNP-based heritability from different functional annotation classes and different cell or tissue types (22) and to examine genetic correlations between the GWAS1, GWAS2, and TIC data sets.
Targeted Replication
The population-based deCODE samples were used 1) to independently replicate the 39 top LD-independent SNPs (r2<0.2 and MAF>0.01; p<1.0×10−5) in the primary meta-analysis, followed by a sign test to examine consistency in the direction of effects in these top SNPs across the two data sets, as well as a targeted meta-analysis of these 39 SNPs using the inverse-variance method (see the Supplemental Methods section); and 2) to examine the genetic relationships between Tourette’s and other tic disorders through polygenic risk score (PRS) analyses (see the Supplemental Methods section) (23). Logistic regressions were performed to test the prediction power of PRS for Tourette’s syndrome and tic disorder case subjects compared with control subjects, adjusted by sex, year of birth, and the first 20 principal components (24).
Polygenic Risk Score Analyses
Genome-wide Tourette’s PRSs adjusted for ancestry principal components (aPRSs) were generated for all subjects in the primary meta-analysis using the entire distribution (GWAS p≤1) of LD-independent SNPs (r2<0.2) through a cross-validation approach (23) and used to examine the relationship between Tourette’s aPRS and ascertainment, family history of Tourette’s or chronic tics, and lifetime worst-ever tic severity (Yale Global Tic Severity Scale total tic score [tic severity], range 0–50) (see the Supplemental Methods section).
Gene-Based and Gene Set Enrichment Analysis
Gene-based tests and competitive gene set enrichment analyses were conducted in MAGMA (25) (see the Supplemental Methods section). Gene-based test statistics were derived using association summary statistics for all SNPs assigned to each gene including 50-kb flanking regions after accounting for LD, and p values were adjusted with a Bonferroni correction for 18,079 genes genome-wide. Gene-based statistics were then analyzed for tissue expression enrichment in 53 distinct human tissues from 714 donors using GTEx RNA-seq data (26), and Bonferroni correction was applied for 53 tissue types (p=0.05/53=9.4×10−4). Tested gene sets included 107 probable autism spectrum disorder susceptibility genes from exome sequencing studies (27), evolutionarily constrained genes (probability of loss-of-function intolerance score >0.9), previously identified constrained genes harboring deleterious rare variants (large copy number variants or de novo loss-of-function mutations) in Tourette’s case subjects (4, 5), and all Gene Ontology terms from the Molecular Signatures Database, version 6.0 (MSigDB 6.0) (see the Supplemental Methods section). Bonferroni correction was applied for the number of gene sets tested.
Results
Genome-Wide Association Study
The final GWAS meta-analysis consisted of 8,265,319 SNPs in 4,819 Tourette’s syndrome case subjects and 9,488 control subjects. No evidence of residual population stratification or systematic technical artifact was observed in any of the individual data sets (see Figure S2 in the online supplement) or in the final meta-analysis (λ=1.072, λ1000=1.011) (Figure 1). LDSC indicated that 86% of the observed test statistic inflation was attributable to an underlying genome-wide polygenic signal (see Figure S3 in the online supplement). PRS analyses in each individual GWAS data set, derived using a leave-one-out approach, as well as in the deCODE sample, indicated genetic homogeneity across all contributing data sets (see Figures S4 and S5 in the online supplement).
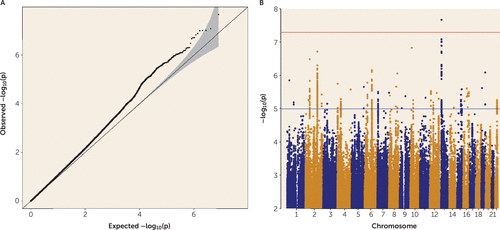
FIGURE 1. Results of the primary Tourette’s syndrome genome-wide association study meta-analysis of 4,819 case subjects and 9,488 control subjectsa
a Panel A is a quantile-quantile plot of observed versus expected −log10(p) values from the primary genome-wide association study (GWAS) meta-analysis. The 95% confidence interval of expected values is indicated in gray. The genomic control λ value is 1.072, and the λ1000 value is 1.011 for single-nucleotide polymorphisms (SNPs) with minor allele frequency >0.01, INFO score (measurement of imputation quality) >0.6, and certainty >0.9. Panel B is a Manhattan plot of all final genotyped and imputed SNPs in the primary Tourette’s syndrome GWAS meta-analysis. The upper horizontal line indicates the genome-wide significance threshold of 5×10−8, and the lower horizontal line indicates the suggestive threshold of 1.0×10−5.
The top SNP in the GWAS meta-analysis, rs2504235, located on chromosome 13q12.2, surpassed the genome-wide significance threshold (odds ratio=1.16, p=2.1×10−8) (Table 1; see also Figure S6 in the online supplement). rs2504235 lies within an intron of FLT3, encoding FMS-like tyrosine kinase 3. No other SNPs achieved genome-wide significance, although rs1933437, a common FLT3 missense variant (Thr227Met) that lies 11.4 kb away from and is in strong LD with rs2504235 (r2=0.93), had a p value of 8.2×10−8 (see Table S3 in the online supplement). Across the genome, 39 LD-independent index SNPs with p values <1×10−5 were identified by LD pruning (r2<0.2) followed by conditional association analyses controlling for the most significant SNP within each 2-Mb window and manual inspection of regional association plots to confirm the presence of supporting statistical evidence of association from nearby SNPs (see Tables S3 and S4 in the online supplement). The top 10 LD-independent index SNPs are presented in Table 1.
| Primary Meta-Analysis | deCODE Sample | Primary and deCODE Sample | ||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| SNP | CHR | BP | A1/A2 | INFO Score | MAF | Odds Ratio | p | MAF | Odds Ratio | p | Odds Ratio | p | LD Block | Genes |
| rs2504235 | 13 | 28,612,886 | A/G | 0.99 | 0.38 | 1.16 | 2.1E–08 | 0.32 | 0.94 | 0.50 | 1.14 | 2.4E–07 | 28591318..28659473 | FLT3 |
| rs191044310 | 10 | 23,705,451 | A/T | 0.83 | 0.02 | 0.54 | 1.5E–07 | 0.0024 | 2.27 | 0.25 | 0.56 | 5.9E–07 | 23661149..23815120 | OTUD1 |
| rs13407215 | 2 | 161,544,891 | T/C | 1.00 | 0.02 | 2.21 | 1.9E–07 | 0.0001 | 0.02 | 0.85 | 2.21 | 1.9E–07 | 160090844..162912453 | Multiple genesc |
| rs2708146 | 2 | 58955953 | G/A | 1.009 | 0.46 | 0.88 | 3.2E–07 | 0.48 | 0.98 | 0.75 | 0.89 | 8.0E–07 | 58847953..59094609 | LINC01122 |
| rs1906252b | 6 | 98,550,289 | A/C | 1.00 | 0.49 | 0.88 | 7.0E–07 | 0.50 | 0.90 | 0.17 | 0.88 | 2.8E–07 | 98214814..98664414 | MIR2113 |
| rs12459560 | 19 | 52,318,380 | T/G | 0.98 | 0.15 | 1.19 | 8.2E–07 | 0.16 | 1.08 | 0.45 | 1.18 | 9.1E–07 | 52266072..52606936 | Multiple genesd |
| rs117648881 | 8 | 113,581,898 | A/G | 0.77 | 0.02 | 0.59 | 8.8E–07 | 0.01 | 0.72 | 0.32 | 0.60 | 6.2E–07 | 113581898..114612903 | CSMD3, MIR2053 |
| rs6670211 | 1 | 29,576,784 | A/C | 1.00 | 0.47 | 0.88 | 1.4E–06 | 0.42 | 0.94 | 0.45 | 0.89 | 1.5E–06 | 29188630..29607279 | EPB41, MECR, OPRD1, PTPRU, SRSF4, TMEM200B |
| rs72853320 | 6 | 36,623,338 | A/G | 1.00 | 0.13 | 1.20 | 1.7E–06 | 0.12 | 0.88 | 0.28 | 1.17 | 2.2E–05 | 36375986..36658092 | CDKN1A, KCTD20, MIR3925, PANDAR, PXT1, RAB44, SRSF3, STK38 |
| rs73205493 | 4 | 2,460,571 | T/C | 0.89 | 0.34 | 1.16 | 1.8E–06 | 0.35 | 1.08 | 0.34 | 1.15 | 1.6E–06 | 2407263..2481088 | LOC402160, RNF4, ZFYVE28 |
TABLE 1. Top 10 linkage disequilibrium–independent loci in the primary Tourette’s syndrome GWAS meta-analysisa
Targeted Replication
The 39 LD-independent index SNPs with p<1×10−5 were investigated for replication in the deCODE sample (706 case subjects, 6,068 control subjects). None of the individual SNPs were replicated after Bonferroni correction (replication threshold for 39 tests, p<0.0013) (see Table 1; see also Table S4 in the online supplement); 23 of 39 putative Tourette’s syndrome risk alleles had the same direction of effect, although this was not statistically significant (binomial two-way sign test, p=0.34).
Meta-analysis restricted to these 39 SNPs was conducted using summary statistics from the primary meta-analysis and the deCODE data with the inverse variance method in METAL. No SNP achieved genome-wide significance; the SNP with the lowest p value was rs13407215, on chromosome 2 (p=1.9×10−7). rs2504235 was not genome-wide significant in this analysis (p=2.4×10−7) (see Table 1; see also Table S4).
Heritability and PRS Analyses
Tourette’s syndrome SNP-based heritability (h2g) was estimated in the primary GWAS meta-analysis using LDSC (h2g=0.21, SE=0.024, p<2.0×10−16). Pairwise genetic correlations across the three independent case-control data sets (GWAS1, GWAS2, TIC) confirmed a significant shared polygenic architecture (GWAS1-GWAS2: rg=0.86, SE=0.21, p=3.9×10−5; GWAS1-TIC: rg=0.84, SE=0.30, p=4.5×10−3; GWAS2-TIC: rg=0.93, SE=0.26, p=4×10−4).
Because the previous estimate of Tourette’s syndrome h2g from the first Tourette’s GWAS (linear mixed model, h2g=0.58, SE=0.09) (7) was significantly higher than that observed in this study, additional heritability analyses were conducted in the individual data sets, stratified on ascertainment status, using linear mixed models (LMM) (7) (Table 2). These analyses confirmed both the high SNP-based heritability of the sibling-pair-enriched Tourette’s GWAS1 sample (GWAS1-LMM: h2g=0.56, SE=0.10; p=1.2×10−9) and the lower heritability of the larger GWAS2 sample (GWAS2-LMM: h2g=0.29, SE=0.04; p=5.5×10−14).
| Case Subjects | Control Subjects | ||||||
|---|---|---|---|---|---|---|---|
| Sample | N | % | N | % | V(G)/Vp_Lb | SE | p |
| GWAS1 | 559 | 14 | 3,400 | 86 | 0.565 | 0.096 | 1.2×10–9 |
| GWAS2c | 2,146 | 46 | 2,564 | 54 | 0.288 | 0.040 | 5.5×10–14 |
| GWAS2c web-based | 934 | 27 | 2,564 | 73 | 0.294 | 0.067 | 2.4×10–6 |
| GWAS2c clinic-based | 1,098 | 30 | 2,564 | 70 | 0.284 | 0.059 | 4.0×10–7 |
TABLE 2. Single-nucleotide polymorphism–based heritability estimates derived using a linear mixed model method for the Tourette’s syndrome GWAS1 and GWAS2 European ancestry case-control samplesa
To explore the hypothesis that the lower heritability of the Tourette’s GWAS2 sample may have arisen from the inclusion of Tourette’s case subjects diagnosed in the community and ascertained using a validated web-based screen (12, 13), the GWAS2 case-control sample was divided into clinic-based and web-based case subjects, and the LMM-based heritability analyses were repeated. Contrary to the predicted hypothesis, both subsets had the same heritability (GWAS2-clinic: h2g=0.29, SE=0.07; p=1.2×10−9; GWAS2-web: h2g=0.28, SE=0.10; p=1.2×10−9) (see Table 2).
Tourette’s Syndrome PRS in Multiplex Versus Simplex Families
Since a large proportion of Tourette’s GWAS1 case subjects were derived from affected sibling-pair families, which might be expected to harbor higher Tourette’s syndrome polygenic risk than case subjects from simplex families without affected first-degree relatives, we examined the relationship between ancestry-adjusted PRS (aPRS) in case subjects from multiplex families (positive for first-degree relative family history) compared with simplex families (negative for first-degree relative family history) (see the Supplemental Methods section).
Because multiplex Tourette’s syndrome case subjects with a Tourette’s-affected parent or sibling (N=417) demonstrated mean aPRSs similar to Tourette’s GWAS case subjects with a chronic tic-affected parent or sibling (N=111) (F=0.12, df=1, p=0.73), we combined both Tourette’s case groups for further analyses (Tourette’s/chronic tic family history positive case subjects, N=528). The combined Tourette’s/chronic tic multiplex case subjects had a significantly higher mean aPRS compared with the aPRS from Tourette’s/chronic tic simplex case subjects (N=346) (F=4.90, df=1, p=0.027), confirming that multiplex case subjects were enriched for Tourette’s polygenic risk (see Figure S7 in the online supplement).
Tourette’s Syndrome PRS and Tic Severity
Given the strong enrichment of Tourette’s aPRS in case subjects from multiplex families, Tourette’s/chronic tic family history positive Tourette’s case subjects were examined next to test whether Tourette’s aPRS may serve as a predictor of higher disease severity in these case subjects (see the Supplemental Methods section). After adjustment for residual population stratification using the first four principal components, higher Tourette’s aPRS was significantly correlated with increased worst-ever tic severity (β=0.93, SE=0.42, p=0.026), with every one-standard-deviation increase in Tourette’s aPRS corresponding to a 0.93-point increase in worst-ever tic severity (total range, 0–50).
Tourette’s Syndrome and Tic Spectrum Phenotypes
Given the hypothesis that Tourette’s and other tic disorders represent a phenotypic spectrum with a shared genetic etiology, Tourette’s PRS derived from the GWAS meta-analysis was compared in Tourette’s and tic spectrum case subjects in the Icelandic deCODE sample (Figure 2; see also Figure S5 in the online supplement). Tourette’s PRS was significantly higher in both deCODE Tourette’s case subjects and tic spectrum case subjects compared with control subjects (odds ratio=1.33, p=5.3×10−9, and odds ratio=1.20, p=5.2×10−4, respectively), explaining 0.78% and 0.42% of the phenotypic variance, respectively. Direct comparison between case groups confirmed that deCODE Tourette’s case subjects carried a higher Tourette’s syndrome polygenic burden than subjects with other tic spectrum disorders (odds ratio=1.14, p=0.05), representing an excess 0.37% of the phenotypic variance (see Figure 2).
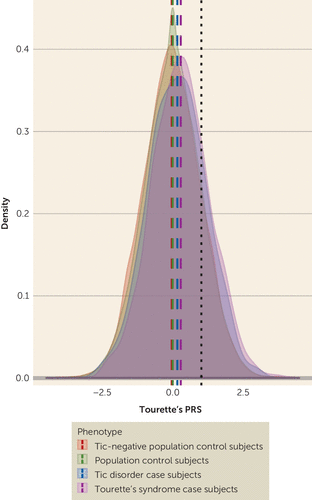
FIGURE 2. Density plot demonstrating the distribution of Tourette’s syndrome polygenic risk scores (PRSs) in population-based Icelandic Tourette’s case subjects, tic disorder case subjects, unscreened population control subjects, and tic-negative control subjectsa
a The x-axis represents the scaled Tourette’s syndrome PRS score, where an increase of one standard deviation in PRS score doubles Tourette’s syndrome risk. The dashed lines correspond to the mean of the respective groups. The black dotted line toward the right corresponds to one standard deviation from the population mean.
Enrichment of Tourette’s Syndrome Heritability by Functional Annotation and Gene Expression
Tourette’s syndrome SNP-based heritability (h2g) from the GWAS meta-analysis was also used as a genome-wide probe to test whether aggregated Tourette’s syndrome genetic risk may be concentrated either in 52 specific functional genomic elements (e.g., promoters, enhancers, epigenetic marks) or in gene expression patterns from 10 grouped tissue or cell types using partitioned LDSC (22). Evolutionarily conserved SNPs (2.6% of all SNPs) were enriched 16.5-fold for Tourette’s h2g, accounting for 42.3% of Tourette’s syndrome heritability (Pr[h2g]/Pr[SNPs]=16.5, SE=5.3, p=3.6×10−3, not significant after correction). A parallel analysis including these evolutionarily conserved SNPs plus 500-bp flanking windows (33% of all SNPs) was enriched 2.8-fold for Tourette’s h2g and accounted for 92.4% of Tourette’s syndrome heritability (Pr[h2g]/Pr[SNPs]=2.80, SE=0.46, p=1.0×10−4; p=0.005 after correction) (see Figure S8 in the online supplement). No other genomic annotations were significantly enriched for Tourette’s SNP-based heritability. In the cell-type analysis, significant enrichment was found only for CNS cell types, with 62.7% of Tourette’s syndrome heritability contributed by 14.8% of SNPs (p=4.2×10−8; p=4.2×10−7 after correction) (see Figure S9 in the online supplement).
Gene-Based Association and Enrichment Analyses
Gene-based association and enrichment tests were performed using meta-analysis summary statistics in MAGMA and gene expression data in GTEx (https://www.gtexportal.org/home/). FLT3 was identified with genome-wide significant association after correcting for 18,079 gene tests (p=8.9×10−7) (see Figure S10 in the online supplement). The most significant SNP in the FLT3 locus, rs2504235, was the only SNP surpassing genome-wide significance threshold in the primary meta-analysis and was significantly associated with FLT3 expression level both in cerebellum (p=6.5×10−10) and cerebral cortex (p=2.6×10−11). No gene set was significantly associated with Tourette’s syndrome after Bonferroni correction. In the gene expression enrichment analyses of 53 adult human tissues, only dorsolateral prefrontal cortex (Brodmann’s area 9) demonstrated significant enrichment of Tourette’s-associated genes after correction (β=0.023, SE=0.0069, p=1.2×10−4) (Figure 3; see also the Supplemental Methods section).
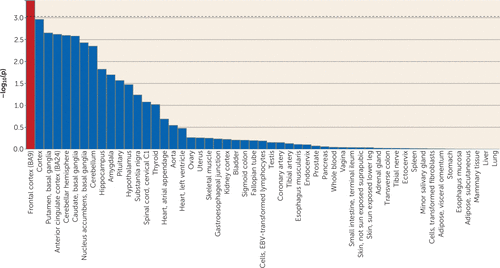
FIGURE 3. Gene expression enrichment analysis of genome-wide Tourette’s syndrome polygenic risk in 53 adult human tissuesa
a Gene-based test statistics were derived from Tourette’s syndrome GWAS meta-analysis summary statistics on single-nucleotide polymorphisms (SNPs) with minor allele frequency >0.01 and INFO score (measurement of imputation quality) >0.9. SNPs were assigned to genes based on their position according to NCBI Build 37.3 and 50-kb upstream and downstream flanking regions. Summary statistics on 18,079 genes were generated. The European panel of the 1000 Genomes data (phase 3) was used as the reference panel to account for linkage disequilibrium. GTEx (version 7) RNA-seq data expression values were log2 transformed with a pseudo-count of 1 after Winsorization at 50, and the average was taken per tissue. Fifty-three specific tissue types were tested separately in MAGMA (23). The significance threshold for the tissue-specific test was calculated using the Bonferroni method (alpha=0.05/53, p<9.43×10−4). Frontal cortex (Brodmann’s area [BA] 9), corresponding to dorsolateral prefrontal cortex, demonstrated significant enrichment of Tourette’s-related genes after correction for multiple hypothesis testing.
Discussion
Tourette’s syndrome has long been conceptualized as part of a spectrum of developmental tic disorders, with transient tics at one end (1) and severe Tourette’s syndrome with multiple psychiatric comorbidities at the other. However, until recently, potential biological relationships between the various tic disorders were unknown, as were the underlying genetic contributions to tic severity. The results of this study further illuminate the genetic architecture of Tourette’s syndrome and its relationships to phenotypic expression. First, the PRS analyses probing the genetic architecture of tic disorders in the population-based Icelandic sample demonstrate that individuals with Tourette’s syndrome share the same underlying polygenic risk as those with other tic disorders. Furthermore, the observation that Tourette’s syndrome case subjects have a significantly higher mean PRS than those with non-Tourette’s tic disorders provides evidence for a liability spectrum of genetic risk within tic disorders. Lastly, within Tourette’s syndrome case subjects, the finding that higher Tourette’s PRS was associated with greater tic severity also builds on our previous analyses demonstrating a relationship between higher Tourette’s PRS and the presence of complex symmetry and socially inappropriate tics (28). These relationships, although hypothesized on the basis of clinical observations, have not previously been demonstrated at the molecular genetic level, and ultimately they will help provide insight into the molecular mechanisms of tic development and expression.
These observations have direct biological and clinical relevance. First, they support previous efforts to conceptualize Tourette’s and chronic tics as a unified condition and to combine them into a single tic spectrum disorder in future diagnostic schemas (1). Although traditionally separated clinically into distinct disorders, chronic/persistent tic disorders, whether consisting of motor tics, vocal tics, or both, appear to be due to the same underlying genetic causes. Second, while the small proportion of explained variance in worst-ever tic severity is a limitation of the present study, work in other polygenic psychiatric disorders, such as schizophrenia, has repeatedly demonstrated that as GWAS sample sizes increase, the proportion of phenotype explained by polygenic risk scores increases markedly (29). It is therefore possible that in the future, Tourette’s PRS may be a potential candidate for predicting both conversion to chronic tics in the 20%−25% of children who present with transient tics (1) and, at the other end of the phenotypic spectrum, tic persistence and lifetime tic severity in those with Tourette’s syndrome. Finally, particularly important in the context of the very large sample sizes required for the success of GWAS efforts, our results suggest that future genetic association studies may benefit from expanding disease definitions to include case subjects with both Tourette’s and chronic tics.
Our genome-wide cell and tissue-based enrichment analyses implicate modulation of gene expression through noncoding variants as a fundamental mechanism in the pathogenesis of Tourette’s syndrome. All of the top tissues in the enrichment analyses were derived from brain, although dorsolateral prefrontal cortex (Brodmann’s area 9) was the only tissue in which eQTL enrichment surpassed Bonferroni correction. The five tissues with the strongest eQTL enrichment (frontal cortex, caudate, putamen, nucleus accumbens, and cerebellum) all represent key nodes within the cortico-striatal and cortico-cerebellar circuits that have been implicated in Tourette’s pathophysiology (1). These results support the hypothesis that Tourette’s syndrome is a developmental circuit disorder affecting motor, cognitive, and behavioral control (as manifested by tics and attention-deficit and obsessive-compulsive symptoms) and suggest that future GWAS analyses in larger data sets should aid in identifying not only the individual genes underlying susceptibility to Tourette’s syndrome but also core pathways in the development and regulation of these circuits that could serve as targets for modulation-based therapies.
Limitations
This study has several potential limitations, the most significant of which is the sample size. Although this is the largest Tourette’s syndrome GWAS conducted to date, our sample of fewer than 5,000 case subjects is clearly not yet sufficient to identify definitive Tourette’s susceptibility variants, as demonstrated by the failure of the top GWAS SNP to replicate in the deCODE sample. Additional potential limitations are also related to sample size, including reduced power to examine additional clinical variables of interest, such as age at onset of tics and co-occurring psychiatric illnesses such as obsessive-compulsive disorder and attention deficit hyperactivity disorder. However, we anticipate that most, if not all, of these limitations can be resolved by substantial increases in the number of Tourette’s syndrome case subjects collected for GWAS, an effort that is currently under way.
1 : Gilles de la Tourette syndrome. Nat Rev Dis Primers 2017; 3:16097Crossref, Medline, Google Scholar
2 : Familial risks of Tourette syndrome and chronic tic disorders: a population-based cohort study. JAMA Psychiatry 2015; 72:787–793Crossref, Medline, Google Scholar
3 : The inheritance of Tourette disorder: a review. J Obsessive Compuls Relat Disord 2014; 3:380–385Crossref, Medline, Google Scholar
4 : Rare copy number variants in NRXN1 and CNTN6 increase risk for Tourette syndrome. Neuron 2017; 94:1101–1111.e7Crossref, Medline, Google Scholar
5 : De novo coding variants are strongly associated with Tourette disorder. Neuron 2017; 94:486–499.e9Crossref, Medline, Google Scholar
6 : De novo sequence and copy number variants are strongly associated with Tourette disorder and implicate cell polarity in pathogenesis. Cell Reports 2018; 24:3441–3454.e12Crossref, Medline, Google Scholar
7 : Partitioning the heritability of Tourette syndrome and obsessive compulsive disorder reveals differences in genetic architecture. PLoS Genet 2013; 9:e1003864Crossref, Medline, Google Scholar
8 : Genome-wide association study identifies five new schizophrenia loci. Nat Genet 2011; 43:969–976Crossref, Medline, Google Scholar
9 : Genome-wide association study of Tourette’s syndrome. Mol Psychiatry 2013; 18:721–728Crossref, Medline, Google Scholar
10 : Genome scan for Tourette disorder in affected-sibling-pair and multigenerational families. Am J Hum Genet 2007; 80:265–272Crossref, Medline, Google Scholar
11 : The Tourette International Collaborative Genetics (TIC Genetics) study, finding the genes causing Tourette syndrome: objectives and methods. Eur Child Adolesc Psychiatry 2015; 24:141–151Crossref, Medline, Google Scholar
12 : Web-based phenotyping for Tourette syndrome: reliability of common co-morbid diagnoses. Psychiatry Res 2015; 228:816–825Crossref, Medline, Google Scholar
13 : Effectiveness of a web-based protocol for the screening and phenotyping of individuals with Tourette syndrome for genetic studies. Am J Med Genet B Neuropsychiatr Genet 2012; 159B:987–996Crossref, Medline, Google Scholar
14 : Second-generation PLINK: rising to the challenge of larger and richer datasets. Gigascience 2015; 4:7Crossref, Medline, Google Scholar
15 : A linear complexity phasing method for thousands of genomes. Nat Methods 2011; 9:179–181Crossref, Medline, Google Scholar
16 : Efficient computation with a linear mixed model on large-scale data sets with applications to genetic studies. Ann Appl Stat 2012; 7:369–390Crossref, Google Scholar
17 : METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics 2010; 26:2190–2191Crossref, Medline, Google Scholar
18 : LD score regression distinguishes confounding from polygenicity in genome-wide association studies. Nat Genet 2015; 47:291–295Crossref, Medline, Google Scholar
19 : Estimation of the multiple testing burden for genomewide association studies of nearly all common variants. Genet Epidemiol 2008; 32:381–385Crossref, Medline, Google Scholar
20 : Estimation of significance thresholds for genomewide association scans. Genet Epidemiol 2008; 32:227–234Crossref, Medline, Google Scholar
21 : Prevalence of tic disorders: a systematic review and meta-analysis. Pediatr Neurol 2012; 47:77–90Crossref, Medline, Google Scholar
22 : Partitioning heritability by functional annotation using genome-wide association summary statistics. Nat Genet 2015; 47:1228–1235Crossref, Medline, Google Scholar
23 : Cross-disorder genome-wide analyses suggest a complex genetic relationship between Tourette’s syndrome and OCD. Am J Psychiatry 2015; 172:82–93Link, Google Scholar
24 : Modeling linkage disequilibrium increases accuracy of polygenic risk scores. Am J Hum Genet 2015; 97:576–592Crossref, Medline, Google Scholar
25 : MAGMA: generalized gene-set analysis of GWAS data. PLOS Comput Biol 2015; 11:e1004219Crossref, Medline, Google Scholar
26 : Genetic effects on gene expression across human tissues. Nature 2017; 550:204–213Crossref, Medline, Google Scholar
27 : Synaptic, transcriptional, and chromatin genes disrupted in autism. Nature 2014; 515:209–215Crossref, Medline, Google Scholar
28 : Identification of two heritable cross-disorder endophenotypes for Tourette syndrome. Am J Psychiatry 2017; 174:387–396Link, Google Scholar
29 : Biological insights from 108 schizophrenia-associated genetic loci. Nature 2014; 511:421–427Crossref, Medline, Google Scholar



