Rare Genome-Wide Copy Number Variation and Expression of Schizophrenia in 22q11.2 Deletion Syndrome
Abstract
Objective:
Chromosome 22q11.2 deletion syndrome (22q11.2DS) is associated with a more than 20-fold increased risk for developing schizophrenia. The aim of this study was to identify additional genetic factors (i.e., “second hits”) that may contribute to schizophrenia expression.
Method:
Through an international consortium, the authors obtained DNA samples from 329 psychiatrically phenotyped subjects with 22q11.2DS. Using a high-resolution microarray platform and established methods to assess copy number variation (CNV), the authors compared the genome-wide burden of rare autosomal CNV, outside of the 22q11.2 deletion region, between two groups: a schizophrenia group and those with no psychotic disorder at age ≥25 years. The authors assessed whether genes overlapped by rare CNVs were overrepresented in functional pathways relevant to schizophrenia.
Results:
Rare CNVs overlapping one or more protein-coding genes revealed significant between-group differences. For rare exonic duplications, six of 19 gene sets tested were enriched in the schizophrenia group; genes associated with abnormal nervous system phenotypes remained significant in a stepwise logistic regression model and showed significant interactions with 22q11.2 deletion region genes in a connectivity analysis. For rare exonic deletions, the schizophrenia group had, on average, more genes overlapped. The additional rare CNVs implicated known (e.g., GRM7, 15q13.3, 16p12.2) and novel schizophrenia risk genes and loci.
Conclusions:
The results suggest that additional rare CNVs overlapping genes outside of the 22q11.2 deletion region contribute to schizophrenia risk in 22q11.2DS, supporting a multigenic hypothesis for schizophrenia. The findings have implications for understanding expression of psychotic illness and herald the importance of whole-genome sequencing to appreciate the overall genomic architecture of schizophrenia.
Chromosomal microarray analysis has made routine the ability to detect pathogenic copy number variations (CNVs), including recurrent CNVs associated with established genomic disorders. Most testing is done postnatally in children with autism spectrum disorder (ASD), developmental delay or intellectual disability, and/or multiple congenital anomalies, or prenatally (1). Several commercial laboratories are now providing the means for noninvasive prenatal screening of microdeletion syndromes (2). With increased early diagnosis, attention is shifting to understanding the later-onset expression of genomic disorders.
Of concern is the association of several genomic disorders with schizophrenia (3–5), a serious psychotic disorder that typically has onset in adolescence or early adulthood and that requires lifelong treatment. The recurrent 22q11.2 deletion associated with 22q11.2 deletion syndrome (22q11.2DS) has an estimated prevalence of one in 3,000–4,000 live births and, in addition to risk for several congenital (e.g., cardiac) anomalies, represents one of the strongest known risk factors for schizophrenia (6). Schizophrenia occurs in about one in four patients with 22q11.2DS, representing a more than 20-fold increase over the general population risk of 1% (6, 7). Given the high but incomplete penetrance of the 22q11.2 deletion for schizophrenia, there is considerable interest in identifying additional genetic factors that may increase the likelihood of an individual developing this illness (6, 8–11).
Multinational collaborative recruitment efforts can facilitate appropriately powered research of individually rare CNVs like 22q11.2 deletions. A goal of the International 22q11.2DS Brain and Behavior Consortium (IBBC) (12) is to discover additional genetic factors that contribute to the high risk for schizophrenia in the 22q11.2DS population. Emerging research suggests that additional rare CNVs elsewhere in the genome may shape the expression of cardiac phenotypes associated with 22q11.2DS (13). In the present study, we used high-resolution genome-wide microarrays and proven CNV detection methods (14) to identify rare CNVs that may be involved in the expression of schizophrenia in 22q11.2DS. We chose this strategy given the relatively high penetrance of rare genic CNVs for idiopathic schizophrenia (15). This is in contrast to common variants, such as single-nucleotide polymorphisms (SNPs), that are located mostly in noncoding regions and that are individually associated with small risks for schizophrenia (15). Data from the IBBC 22q11.2DS cohort enabled us to compare genome-wide burden and genic content of rare autosomal CNVs, outside of the 22q11.2 deletion region, between individuals with schizophrenia or related psychotic disorders and those with no psychotic disorder at age ≥25 years.
Method
Ascertainment of Samples From 22q11.2DS Subjects for CNV Data
Subjects with a presumed 22q11.2 deletion were recruited from 22 international sites (see Table S1 in the data supplement that accompanies the online edition of this article) and provided DNA samples that were genotyped using the Affymetrix Genome-Wide Human SNP Array 6.0 at the Albert Einstein College of Medicine (Bronx, N.Y.) (13). Informed consent was obtained from all subjects and/or their legal guardian. This study was approved by the local institutional research ethics boards of each site.
Similar to our previous studies of CNV (14), all downstream processing and analysis of microarray data, including applying rigorous methods for quality control and CNV detection, were completed at the Centre for Applied Genomics in Toronto. Initially, there were 866 DNA samples available from subjects with a presumed 22q11.2 deletion and with psychiatric phenotype data. Of these, 741 (85.6%) samples had high-quality CNV data after accounting for batch effects and applying other quality-control measures to delineate an unrelated data set of similar ancestry (N=666; see the online data supplement for full details).
Phenotype and Diagnostic Group Determination
Of the 666 unrelated 22q11.2DS subjects of European ancestry with genome-wide CNV data, 329 met inclusion criteria for this study. The phenotypic and diagnostic assessment protocol is described elsewhere (12). We first selected subjects who met DSM-IV-TR diagnostic criteria for a major psychotic disorder; 15 subjects with an affective psychosis were excluded. The resulting 158 individuals formed the “schizophrenia group” and had specific diagnoses of schizophrenia (N=117; 74.1%); schizoaffective disorder (N=12; 7.6%); psychosis not otherwise specified (N=20; 12.7%); or schizophreniform (N=1; 0.6%), delusional (N=2; 1.3%), brief psychotic (N=2; 1.3%), or other psychotic (N=4; 2.5%) disorders. Median age at onset of psychotic illness was 21 (range=7–59) years, with no overall sex difference. We assigned individuals to the “nonpsychotic group” (N=171) if they had no history of any psychotic illness when assessed at age ≥25 years (median age=32, range=25–67). As expected for 22q11.2DS, given the known impact of ascertainment and disease factors (6), in the nonpsychotic group compared with the schizophrenia group, the proportion of female subjects was higher (N=113, 66.1%; N=82, 51.9%, respectively; p=0.0089), the proportion of cases with a congenital cardiac defect where known (N=248) was higher (N=74, 58.3%; N=49, 40.5%, respectively; p=0.0051), and the median IQ was higher (75, range=34–105 [N=137]; 69, range=36–94 [N=121], respectively; p<0.0001).
CNV Detection and Annotation
Similar to previous studies (14), we used CNV methods that ensure high (>90%) validation using a second laboratory method (16) (see Table S2 in the online data supplement). Genome-wide CNVs were identified with a multiple-algorithm approach using Birdsuite (17), iPattern (18), and Affymetrix Genotyping Console (http://www.affymetrix.com) to maximize CNV call accuracy. We included CNVs for analyses only if they were identified by two or three of the CNV calling algorithms, if they spanned 10 consecutive array probes for deletions or duplications, and if there was <75% overlap with sequence from segmental duplications. We used 9,611 independent population-based controls of European ancestry (see Table S2 in the data supplement) to adjudicate CNV rarity. Using a stringent cutoff, we designated CNVs found in <0.1% of the population controls as “rare” in the 22q11.2DS subjects. The CNVs were adjudicated for microRNA (miRNA) content using miRBase (19) and for genic content using RefSeq. The CNVs were deemed exonic if they overlapped at least one base pair (bp) of coding sequence. All genomic coordinates refer to GRCh37/hg19.
CNV Burden Analysis and Statistical Approach
For CNV burden analyses, we compared the proportion of subjects with one or more rare autosomal CNVs ≥10 kb (all, deletions, duplications, and restricting to those overlapping a gene, exon, or miRNA) between the schizophrenia and nonpsychotic groups. Fisher’s exact tests, odds ratios, and 95% confidence intervals (CIs) were calculated using SAS (SAS Institute, Cary, N.C.) or R 3.3.1 software. All tests were two-sided, with p<0.05 defined for statistical significance.
Gene Set Enrichment Analysis
To assess whether genes overlapped by rare CNVs in the schizophrenia group were overrepresented in functional pathways related to schizophrenia, we evaluated 19 gene sets (see Table S3 in the data supplement) similar to those used in a recent schizophrenia consortium study (16). These comprised genes with roles in human neurodevelopment, neuronal function, or synaptic function (seven sets), or human orthologs of mouse genes whose disruptions cause neurobehavioral or nervous system abnormalities (three sets). Given evidence for miRNA mechanisms in schizophrenia and the role of DGCR8 in genome-wide miRNA buffering (9, 20), we also examined genes predicted to be targets of miRNAs showing differential gene expression with haploinsufficiency of DGCR8 in a mouse model (one set) (9, 10, 20). Eight further gene sets involved genes associated with abnormal phenotypes in nonbrain organ systems (seven sets), or with causing prenatal or perinatal lethality (one set), in mice (16).
The gene set burden analysis used a logistic regression deviance test (21) (R/Bioconductor package cnvGSA: Gene Set Analysis of (Rare) Copy Number Variants [version 1.18.0]) (https://www.bioconductor.org/packages/release/bioc/html/cnvGSA.html) to evaluate if the number of genes overlapped by rare exonic deletions or duplications in each subject for each gene set (i.e., gene set–specific genic burden) is predictive of the subject being a member of the schizophrenia or nonpsychotic group. We report the regression coefficients after standardizing the gene set gene count. Sex was used as a covariate. Similar to our previous whole-genome sequencing study of 22q11.2DS (9), we also performed a secondary gene set burden analysis that included each original gene set after restricting to only the genes that are predicted to be targets of DGCR8 (9). Multiple-testing correction (the Benjamini-Hochberg false discovery rate) was performed separately for each gene set group (with and without DGCR8 restriction) and CNV type (deletions, duplications). Gene sets with a Benjamini-Hochberg false discovery rate <10% and a p value <0.05 were considered to be significantly enriched (9). To account for overlap between the gene sets, we retested the burden following a stepwise logistic regression approach that used the same regression deviance test and covariates after ranking the gene sets based on most to least significant p values.
Network Connectivity Test and Network Construction
We performed an absolute connectivity test by treating the 46 protein-coding genes (N=42 with network data) in the 22q11.2 deletion region as “bait” and the 41 genes from the mouse abnormal nervous system gene set overlapped by duplications in the 22q11.2DS schizophrenia group (N=39 with network data) as “prey.” For each bait gene, the top 200 interaction neighbors (prey) were retrieved using GeneMANIA physical protein-protein interaction and pathway interaction networks (22). Because each network contains different information, each requires weighting based on its capability to predict gene functions as defined by Gene Ontology (22). Accordingly, we used the weighting procedure implemented in GeneMANIA and selected Gene Ontology biological process functional annotations to preweight the selected protein interaction networks (22) as more representative of pathways than Gene Ontology cellular component and molecular function. We determined a connectivity score for each bait gene using the average of the respective preys’ GeneMANIA scores and performed an empirical test to compare this connectivity score with that for the 15,824 genes outside the 22q11.2 deletion region. Nominal p values were then corrected using the Benjamini-Hochberg false discovery rate. We generated a gene interaction proximity network using the results from the network connectivity analysis, as well as “linker” genes identified using GeneMANIA (22), visualizing the resulting network in Cytoscape (23).
Results
22q11.2 Deletions
There was no significant between-group difference in the distribution of 22q11.2 deletions (χ2=0.18, df=1, p=0.68). The majority of subjects had LCR22A–LCR22D 22q11.2 deletions (N of schizophrenia subjects, 147, 93.0%; N of nonpsychotic subjects, 157, 91.8%; see Figure S1 in the online data supplement). Three subjects (1.9%) in the schizophrenia group had a known proximal variant of the LCR22A–LCR22D deletion, and the remainder had an LCR22A–LCR22B (N of schizophrenia subjects, 7, 4.4%; N of nonpsychotic subjects, 11, 6.4%) or an LCR22A–LCR22C (N of schizophrenia subjects, 1, 0.6%; N of nonpsychotic subjects, 3, 1.8%) deletion (see Figure S1 in the data supplement).
Quantitative Genome-Wide Burden of Additional Rare CNVs
In the overall sample of 329 subjects with a 22q11.2 deletion, there were 726 additional rare genome-wide autosomal CNVs in 291 (88.4%) subjects (N of schizophrenia subjects, 142, 89.9%; N of nonpsychotic subjects, 149, 87.1%; see Tables S4 and S5 in the data supplement). The proportion of schizophrenia subjects (N=16, 10.1%) with one or more rare CNVs overlapping an miRNA was not significantly greater than that of the nonpsychotic group (N=8, 4.7%; p=0.0878).
There were 272 rare CNVs (173 duplications, 99 deletions) that overlapped at least one bp of coding sequence in 182 (55.3%) subjects. Those with exonic duplications (N of schizophrenia subjects, 68, 43.0%; N of nonpsychotic subjects, 59, 34.5%) outnumbered those with exonic deletions (N of schizophrenia subjects, 38, 24.1%; N of nonpsychotic subjects, 47, 27.5%) in both subject groups. There were no overall between-group differences in the total number of rare CNVs (deletions and/or duplications; see Table S4 in the data supplement) or in the total genomic length of rare CNVs per subject (median=173 kb and range=12 kb–3.6 Mb in the schizophrenia group, compared with median=184 kb and range=11 kb−1.9 Mb in the nonpsychotic group; p=0.48). Restricting to rare exonic CNVs showed similar results (data not shown).
However, when examining the total number of genes (regardless of function) overlapped by the additional rare CNVs, there were significantly more genes overlapped by rare exonic deletions in the schizophrenia group (median=2 genes, range=1–18) compared with the nonpsychotic group (median=1 gene, range=1–14) (p=0.0058) (Table 1). There was no such significant finding for duplications. There was no significant difference in the IQ of individuals with and without an additional rare exonic deletion in either the schizophrenia group or the nonpsychotic group (data not shown). Multigenic rare deletions in the schizophrenia group included a 410 kb 1q21.1 deletion (at the TAR locus) and a 600 kb 16p12.2 deletion, both previously associated with schizophrenia in general population samples (3).
| Variable | 22q11.2DS-Schizophrenia (N=142)a | 22q11.2DS-Nonpsychotic (N=149)a | Analysis | |||
|---|---|---|---|---|---|---|
| Median | Range | Median | Range | z | p | |
| Deletion or duplication | 2 | 1–26 | 2 | 1–26 | 1.5787 | 0.1144 |
| Deletionb | 2 | 1–18 | 1 | 1–14 | 2.7569 | 0.0058 |
| Duplication | 2 | 1–26 | 2 | 1–26 | 0.1470 | 0.8831 |
TABLE 1. Average Number of Genes Overlapped by Additional Rare Exonic Autosomal CNVs in 22q11.2DS
Functional Burden of Additional Rare CNVs
We performed a gene set enrichment analysis to identify between-group differences in the burden of rare CNVs overlapping functionally related genes (i.e., gene sets or pathways). Compared with the nonpsychotic group, after multiple-test correction (Benjamini-Hochberg false discovery rate <10% and nominal p value <0.05), the schizophrenia group had significantly more individuals with rare exonic duplications overlapping genes from six of the 19 gene sets assessed (Table 2). There were no between-group differences for deletions. Of the six significantly enriched gene sets, only the top-ranking nervous system phenotype gene set remained significant (p=0.00062) in the stepwise logistic regression model (Table 2); overlap of the gene signal with the other five gene sets is shown in Table S6 in the online data supplement. Secondary gene set analyses, restricted to genes predicted to be affected by DGCR8 haploinsufficiency (9, 20), showed significant improvement in the burden analysis results beyond that for the full gene set only for the muscle/cardiovascular gene set, after permutation to account for gene set size (data not shown).
| 22q11.2DS-Schizophrenia (N=142)b | 22q11.2DS-Nonpsychotic (N=149)b | Analyses | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Gene Set Name | Total Genes (N) | CNVs (N) | Subjects | CNVs (N) | Subjects | p | FDR-BH | Odds Ratio | ||
| N | % | N | % | |||||||
| Nervous system phenotype (MGI) | 2,609 | 37 | 32 | 22.5 | 16 | 15 | 10.1 | 0.00062 | 0.012 | 2.36 |
| Neurobehavioral phenotype (MGI) | 2,602 | 37 | 33 | 23.2 | 19 | 19 | 12.8 | 0.00174 | 0.016 | 2.21 |
| Neural phenotype union (MGI) | 3,764 | 48 | 42 | 29.6 | 25 | 24 | 16.1 | 0.00260 | 0.016 | 1.80 |
| Muscle/cardiovascular phenotype (MGI) | 2,327 | 27 | 25 | 17.6 | 13 | 13 | 8.7 | 0.00481 | 0.022 | 2.38 |
| Endocrine/exocrine/reproduction phenotype (MGI) | 2,298 | 31 | 31 | 21.8 | 18 | 17 | 11.4 | 0.01099 | 0.042 | 2.08 |
| Synaptic (Gene Ontology) | 860 | 15 | 14 | 9.9 | 6 | 6 | 4.0 | 0.02472 | 0.078 | 2.81 |
TABLE 2. Gene Sets Showing Significant Enrichment in the 22q11.2DS-Schizophrenia Group for Rare Exonic Duplicationsa
In 32 subjects with 22q11.2DS-schizophrenia, there were 37 rare duplications that contributed to the nervous system phenotype gene set result (Table 2 and Table 3). For individuals with IQ data available, there was no significant difference in the median IQ between those with (N=23) and without (N=104) a rare duplication contributing to this gene set result (68 compared with 70, p=0.41). Four (10.8%) duplications were recurrent, flanked by segmental duplications, and previously associated with idiopathic schizophrenia (Table 3). These included two 22q11.2DS-schizophrenia subjects with 1.9–2.0 Mb duplications at 15q13.2-q13.3 (BP4-BP5) (26), one with a 1.5 Mb duplication at 17q12 (27), and one with a 1.3 Mb duplication at 22q11.23 (28). Eighteen (48.6%) of the 37 CNVs overlapped a single gene, including several genes implicated in idiopathic schizophrenia by rare variant studies, such as RYR2 (3, 21), GRM7 (3, 29), DOK7 (3, 30), and CACNA2D1 (3, 31). In silico breakpoint analysis indicated that the majority (N=28; 65.1%) of the genes implicated in the rare duplication gene set results had coding sequence that was disrupted by the CNV (data not shown).
| CNV | Subject | Chromosome | Cytoband | Startb | Size | Genes (N) | Very Rare CNVc | Flanking Low-Copy Repeats | All Genes Overlapped by CNV | Gene(s) Contributing to Nervous System Gene Set Results |
|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 17 | 1 | 1p22.1 | 94463617 | 18760 | 1 | • | ABCA4 | ABCA4 | |
| 2 | 130 | 1 | 1q32.2 | 209881045 | 28340 | 1 | • | HSD11B1 | HSD11B1 | |
| 3 | 93 | 1 | 1q43 | 237492310 | 26855 | 1 | • | RYR2 | RYR2 | |
| 4 | 144 | 2 | 2q35 | 216834680 | 340811 | 5 | • | TMEM169, MARCH4, PECR, XRCC5, MREG | XRCC5 | |
| 5 | 39 | 3 | 3p26.1 | 7619772 | 536651 | 1 | • | GRM7 | GRM7 | |
| 6 | 55 | 3 | 3p22.2 | 38608585 | 510439 | 4 | • | WDR48, SCN11A, SCN5A, SCN10A | SCN11A, SCN5A, SCN10A | |
| 7 | 28 | 3 | 3q29 | 195263161 | 232072 | 4 | APOD, MUC4, MUC20, PPP1R2, miR–570–5p, miR–570–3p | APOD | ||
| 8 | 130 | 4 | 4p16.3 | 3463452 | 10187 | 1 | • | DOK7 | DOK7 | |
| 9 | 42 | 4 | 4p15.2 | 26433603 | 49693 | 2 | • | RBPJ, CCKAR | CCKAR | |
| 10 | 65 | 4 | 4q35.1 | 186979699 | 76584 | 1 | TLR3 | TLR3 | ||
| 11 | 110 | 6 | 6q11.1 | 62234532 | 270027 | 2 | MTRNR2L9, KHDRBS2 | KHDRBS2 | ||
| 12 | 15 | 6 | 6q26 | 161687430 | 1202784 | 2 | • | AGPAT4, PARK2 | PARK2 | |
| 13 | 150 | 6 | 6q27 | 168167050 | 228482 | 2 | • | MLLT4, HGC6.3 | MLLT4 | |
| 14 | 141 | 7 | 7p21.2 | 13946099 | 33357 | 1 | • | ETV1 | ETV1 | |
| 15 | 55 | 7 | 7q21.11 | 81993260 | 185718 | 1 | CACNA2D1 | CACNA2D1 | ||
| 16 | 114 | 8 | 8p23.3 | 427857 | 77504 | 1 | • | TDRP | TDRP | |
| 17 | 127 | 8 | 8p22 | 13329776 | 290648 | 2 | • | DLC1, C8orf48 | DLC1 | |
| 18 | 110 | 8 | 8p12 | 34906448 | 250455 | 1 | • | UNC5D | UNC5D | |
| 19d | 143 | 8 | 8q12.1 | 56722133 | 159268 | 2 | LYN, TGS1 | LYN | ||
| 20 | 79 | 9 | 9q22.2 | 93616404 | 82699 | 1 | SYK | SYK | ||
| 21 | 10 | 10 | 10q23.31 | 89651696 | 46930 | 1 | • | PTEN | PTEN | |
| 22 | 35 | 10 | 10q24.2 | 99411165 | 39708 | 2 | AVPI1, PI4K2A | PI4K2A | ||
| 23 | 15 | 11 | 11q13.5 | 76735763 | 156340 | 4 | • | B3GNT6, OMP, MYO7A, CAPN5 | MYO7A, OMP | |
| 24 | 86 | 11 | 11q23.3 | 115010867 | 139977 | 1 | • | CADM1 | CADM1 | |
| 25 | 12 | 12 | 12q24.21, 12q24.22 | 116396113 | 1021914 | 6 | • | FBXW8, MAP1LC3B2, MED13L, RNFT2, C12orf49, HRK, miR–4472, miR–620 | FBXW8 | |
| 26 | 116 | 13 | 13q12.12 | 23397621 | 1690189 | 9 | • | SPATA13, C1QTNF9, C1QTNF9B, SGCG, C1Q, TNF9B-AS1, PARP4, MIPEP, SACS, TNFRSF19, miR–2276–5p, miR–2276–3p | C1QTNF9B, TNFRSF19 | |
| 27 | 30 | 14 | 14q32.11 | 91288380 | 77382 | 1 | • | RPS6KA5 | RPS6KA5 | |
| 28e | 92 | 15 | 15q13.2,15q13.3 | 30668479 | 1952748 | 9 | • | KLF13, OTUD7A, MTMR10, FAN1, CHRNA7, LOC283710, ARHGAP11B, TRPM1, CHRFAM7A, miR–211–5p, miR–211–3p | CHRNA7, TRPM1 | |
| 29e | 52 | 15 | 15q13.2,15q13.3 | 30747394 | 2129590 | 9 | • | KLF13, OTUD7A, MTMR10, FAN1, GOLGA8O, CHRNA7, LOC283710, ARHGAP11B, TRPM1, miR–211–5p, miR–211–3p | CHRNA7, TRPM1 | |
| 30 | 131 | 15 | 15q25.1 | 78421669 | 15345 | 1 | • | CIB2 | CIB2 | |
| 31 | 33 | 16 | 16q24.3 | 89420230 | 258822 | 4 | • | RPL13, SPG7, CPNE7, ANKRD11 | SPG7 | |
| 32d,e | 132 | 17 | 17q12 | 34816256 | 1428102 | 15 | • | ZNHIT3, LHX1, DUSP14, MRM1, ACACA, DDX52, DHRS11, SYNRG, C17orf78, HNF1B, AATF, MYO19, PIGW, TADA2A, GGNBP2, miR–378j, miR–2909 | LHX1 | |
| 33 | 147 | 18 | 18q23 | 74711326 | 80540 | 1 | • | MBP | MBP | |
| 34 | 34 | 20 | 20p13 | 4895803 | 181274 | 1 | • | SLC23A2 | SLC23A2 | |
| 35 | 130 | 20 | 20p11.21 | 25061344 | 198784 | 3 | • | ENTPD6, PYGB, VSX1 | VSX1 | |
| 36 | 41 | 21 | 21q22.2 | 41337036 | 120608 | 1 | • | DSCAM | DSCAM | |
| 37 | 121 | 22 | 22q11.23 | 23675258 | 1384582 | 26 | • | SMARCB1, MMP11, C22orf43, DDTL, RGL4, SLC2A11, MIF, CHCHD10, DERL3, C22orf15, IGLL1, GSTT2B, ZNF70, DDT, VPREB3, GSTT2, ADORA2A, SNRPD3, FAM211B, CABIN1, SPECC1L, GUCD1, SUSD2, GGT5, UPB1, GGT1 | ADORA2A, MIF, SMARCB1 |
TABLE 3. Genes Contributing to the Nervous System Gene Set Results for Rare Genome-Wide Exonic Duplications Showing Significant Enrichment in the 22q11.2DS-Schizophrenia Groupa
Potential Mechanisms of Rare CNV-Related Risk for Schizophrenia in 22q11.2DS
We attempted to identify potential mechanisms contributing to the schizophrenia risk posed by the additional rare duplications overlapping genes from the nervous system phenotype gene set (Table 3). Using protein-protein interaction data available from GeneMANIA (22), we empirically tested whether genes in the 22q11.2 region (N=46 total, 42 with network data) were more connected to the nervous system phenotype genes overlapped by rare duplications in the schizophrenia group (N=41 total, 39 with network data) when compared with 15,824 other protein-coding genes in the genome. This analysis revealed four 22q11.2 deletion region genes with significant interactions at a false discovery rate <20% (nominal p value <0.02): P2RX6, SEPT5, RTN4R, and CLTCL1 (see Table S7 in the data supplement; Figure 1). The a priori probability of randomly extracting 42 genes from the genome with at least four genes achieving significant connectivity at a false discovery rate <20% is <0.0098, providing further support for this result.
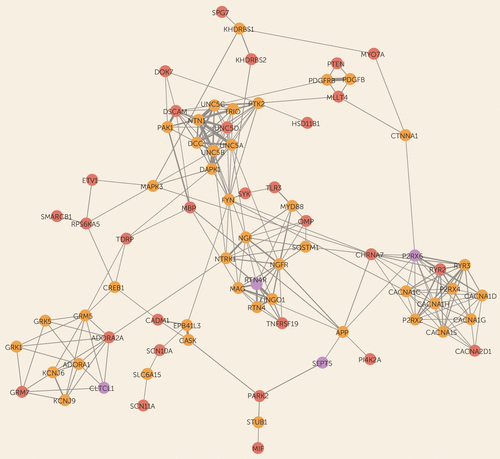
FIGURE 1. Gene Interaction Proximity Network Based on Physical and Pathway Interactionsa
a Circles correspond to genes: 22q11.2 deletion genes with significant connectivity (CLTCL1, P2RX6, RTN4R, and SEPT5; purple), genes with mouse abnormal nervous system phenotypes overlapped by duplications exclusively in the 22q11.2 deletion syndrome–schizophrenia group (red), and linker genes (gold). The initial sets comprised the four 22q11.2 deletion region genes with significant connectivity and the 39 genes overlapped by duplications found exclusively in schizophrenia subjects and within the nervous system phenotype gene set. Important “linker” genes were identified by retrieving the top 25 interaction neighbors of these genes in GeneMANIA, using the same protein-protein interaction and pathway interaction networks used in the connectivity test, and then subsetting to those found connected to at least two different bait genes and belonging to the nervous system phenotype gene set (86 genes). Connections correspond to interaction proximity obtained from GeneMANIA, setting the limit to the top 50 interactors; the connection line thickness is proportional to the GeneMANIA score (22). The resulting network is visualized in Cytoscape (23). The network suggests the presence of four interaction clusters, characterized by the presence of one gene from the 22q11.2 deletion region with significant connectivity results and at least one schizophrenia-only duplication-related gene (see text for details).
The interaction proximity network constructed using the four 22q11.2 deletion region genes and the genes implicated by the duplication gene set findings suggested a diverse set of pathways potentially related to schizophrenia pathogenesis (Figure 1). Four interaction clusters were characterized by the presence of a significant gene from the 22q11.2 deletion region and/or at least one schizophrenia-only gain gene: axon guidance and neuronal adhesion (UNC5D, DSCAM), axon guidance and nerve growth factor signaling (RTN4R; TNFRSF19), purinergic receptors and calcium channels (P2RX6; CACNA2D1), and glutamatergic and adenosine receptors, synaptic trafficking (CLTCL1; ADORA2A, GRM7) (Figure 1).
Discussion
The results of the present study provide new data relevant to a key knowledge gap in our understanding of the association between genomic disorders like 22q11.2DS and a profoundly increased risk for developing major psychotic illnesses such as schizophrenia. For the first time, there are sufficient data to support additional rare CNVs that overlap protein-coding sequence as contributing factors to this increased risk. These comprise both rare duplications that overlap genes involved in neuronal function and rare deletions that tend to overlap more genes in those who develop schizophrenia. The findings suggest the possibility that additional information captured by chromosomal microarray at the time of initial diagnosis of a 22q11.2 deletion could eventually help in identifying some of those at greatest risk for schizophrenia.
The results are consistent with emerging research suggesting that rare deleterious variants elsewhere in the genome, in addition to the 22q11.2 deletion, are likely to be involved in the variability of neuropsychiatric and other developmental expression of this microdeletion syndrome (6, 8–11). This finding has potential implications for the schizophrenia risk associated with the CNVs of other genomic disorders, such as the 3q29 deletion (32) or the 16p11.2 duplication (33). The results are also in keeping with data from the general population of individuals with neurodevelopmental disorders, where approximately 4% have two rare contributing genetic mutations or genetic diagnoses (34). Although there are case reports of two rare CNVs in patients with schizophrenia (35, 36), there are few previous studies of genome-wide CNVs in patients with typical 22q11.2 deletions, and these involved small samples and/or used low resolution arrays or various methods to assess CNVs and their rarity (8, 35–37). Only two of these small studies compared schizophrenia and nonpsychotic groups within 22q11.2DS (8, 37).
The results of this study are in line with findings supporting an association between large, usually multigenic, rare duplications and schizophrenia in the general population (5, 14). Similarly, the results for enriched burden of genes overlapped by rare deletions in 22q11.2DS-schizophrenia also appear consistent with studies of schizophrenia in the general population (16) and with a recent rare CNV study reporting a higher number of overlapped genes in a large schizophrenia cohort compared with controls (38). Within 22q11.2DS, the structural variant results in the present study complement those of pilot whole-genome sequencing findings for 22q11.2DS using an extreme phenotype approach to study additional genome-wide rare sequence variants affecting schizophrenia expression (9). In that pilot study, there was a significantly enriched burden of brain function–related genes with high-quality nonsynonymous deleterious missense and loss of function sequence variants in the schizophrenia cases (9). Genes involved included RYR2 (3, 39) and BSN (40), the latter encoding a presynaptic scaffolding protein, bassoon (9). Results of the present study (Table 3) further implicate these novel genes and pathways for schizophrenia. The RYR2 gene and two genes (DLC1 and ABCA4) encoding members of a BSN superfamily of neuronal transmembrane proteins that regulate neurotransmitter release at the synapse were overlapped by rare duplications in subjects with 22q11.2DS and schizophrenia. BSN was also implicated in the synaptic network derived from a large CNV study of schizophrenia (16), as were some of the network connections in the present study (Figure 1).
The significant genetic interactions between 22q11.2 deletion region genes and duplication locus genes implicate other pathways of potential interest to schizophrenia pathogenesis. These include the RTN4R gene encoding the Nogo receptor and its coreceptor, TNFRSF19, involved in mediating axonal growth inhibition, regeneration, and plasticity and in limiting the excitatory synapse number during brain development (41). Similarly, the P2RX6-related functional gene cluster includes RYR2, CHRNA7, and CACNA1C, previously implicated in schizophrenia pathogenesis (42). Several of the genes in the network (Figure 1) have also been implicated as targets of 22q11.2 region miRNAs or of miRNAs dysregulated by DGCR8 haploinsufficiency (10). Interestingly, there is also some evidence from a recent brain expression study implicating RTN4R and P2RX6 down-regulation in the pathogenesis of idiopathic schizophrenia (43), further supporting the potential generalizability of the findings in the present study.
The 22q11.2 deletion may lower the threshold for expression of the multiple mechanisms and/or pathways involved in causing schizophrenia, within and between individuals (9, 10). The results of the present study primarily implicate additional genome-wide rare CNVs that affect the protein-coding sequence of genes in the expression of schizophrenia in the context of a 22q11.2 deletion. Effect sizes may be smaller for other mechanisms. For example, noncoding regulatory regions and/or miRNAs (at the trend level in this study) may also play important roles in schizophrenia, including in 22q11.2DS, as suggested by rare sequence variant findings for 22q11.2DS supporting DGCR8 haploinsufficiency and altered miRNA buffering mechanisms (9, 20). Power may also be an issue for the difference observed in additional rare CNV findings in 22q11.2DS between those for duplications and deletions. A more neuro-specific effect for deletions could perhaps become apparent if the sample size were larger. However, mechanisms for expression of deletions and duplications may differ (1), and penetrance is, on average, higher for deletions than for duplications (35). In the presence of the 22q11.2 deletion, additional rare deletions may be more likely to potentially implicate pleiotropic genes in expression of schizophrenia, whereas duplications could require genes having greater specific impact on neuronal functioning. Notably, the rare duplication burden in 22q11.2DS-schizophrenia was enriched for genes associated with mouse neurobehavioral phenotypes, similar to results in a recent large case-control study of schizophrenia, although the latter involved somewhat smaller effect sizes (odds ratios of 1.18 and 1.68, respectively) and deletions, including most commonly the 22q11.2 deletion (16). Results from the present study provide some evidence for the expression of schizophrenia in 22q11.2DS involving network interactions that include genes in the 22q11.2 deletion region and genes overlapped by other rare structural variants outside this locus.
Advantages and Limitations
Despite having the largest sample of 22q11.2DS subjects with and without a psychotic disorder to study the impact of additional rare CNVs on expression, power was limited by the available cohort size. However, the signal related to schizophrenia expression imparted by additional rare genome-wide CNVs produced significant results. This is remarkable, given the virtually ubiquitous presence of any neurodevelopmental phenotype in 22q11.2DS (44) and the likelihood that some subjects in the nonpsychotic group may go on to develop schizophrenia. It is expected that some of the rare CNVs associated with schizophrenia and the genes implicated may also be involved in expression of other neurodevelopmental disorders, consistent with the literature supporting overlap of pathways (15). However, the results do not support intellect as a primary driver of the findings, and while most subjects were not systematically assessed for the presence of ASD, this was documented for few subjects (N=18; 5.5%). The ability to recruit more adults with no psychotic illness and individuals with schizophrenia within the 22q11.2DS population could provide the means to identify the strength of the relationship between each additional CNV and the risk for schizophrenia, as well as other mechanisms for schizophrenia expression where the effect sizes are lower. In future studies, larger samples of 22q11.2DS may also assist in determining the role of X chromosome variants and whether, as in ASD and intellectual disability (45), the X chromosome may provide some protective capacity for schizophrenia expression in the context of a 22q11.2 deletion.
The IBBC proband–only 22q11.2DS sample provides no capability of examining inheritance, parent-of-origin, or segregation data for the 22q11.2 deletion or the additional CNVs, or the family history of schizophrenia, all of which are factors of interest in disease expression (11). In smaller samples, the parental origin and inheritance status of the 22q11.2 deletion have not appeared to play a role in schizophrenia expression, however (8), and one can expect that >90% of the 22q11.2 deletions would be de novo mutations (6).
The results suggest that additional rare CNVs overlapping genes outside of the 22q11.2 deletion region contribute to schizophrenia risk in 22q11.2DS. The fact that these results involved fewer than half of the subjects with schizophrenia supports the likelihood that other unidentified variants and mechanisms also play a role in expression. The recurrent de novo 22q11.2 deletion has no relationship to parental age and can derive from either parental chromosome (6, 8), but the inheritance of additional genome-wide CNVs remains to be determined. The findings support a multigenic hypothesis for schizophrenia and further illustrate the value of a 22q11.2DS model for identifying genome-wide rare variants to help delineate the genetic architecture of idiopathic schizophrenia. Lessons learned from 22q11.2DS may be applicable to other recurrent CNVs that contribute to risk for schizophrenia in the general population (14). Whole-genome sequencing will allow for more detailed assessment of CNV breakpoints, as well as the overall genetic architecture and proposed mechanisms of schizophrenia (9, 20).
1 : Consensus statement: chromosomal microarray is a first-tier clinical diagnostic test for individuals with developmental disabilities or congenital anomalies. Am J Hum Genet 2010; 86:749–764Crossref, Medline, Google Scholar
2 : Clinical experience with single-nucleotide polymorphism-based non-invasive prenatal screening for 22q11.2 deletion syndrome. Ultrasound Obstet Gynecol 2016; 47:177–183Crossref, Medline, Google Scholar
3 : Prenatal genetic testing with chromosomal microarray analysis identifies major risk variants for schizophrenia and other later-onset disorders (letter). Am J Psychiatry 2013; 170:1498Link, Google Scholar
4 : Delineating the 15q13.3 microdeletion phenotype: a case series and comprehensive review of the literature. Genet Med 2015; 17:149–157Crossref, Medline, Google Scholar
5 : Parental origin of interstitial duplications at 15q11.2-q13.3 in schizophrenia and neurodevelopmental disorders. PLoS Genet 2016; 12:e1005993Crossref, Medline, Google Scholar
6 : 22q11.2 deletion syndrome. Nat Rev Dis Prim 2015; 19:15071Crossref, Google Scholar
7 : Elevated prevalence of generalized anxiety disorder in adults with 22q11.2 deletion syndrome (letter). Am J Psychiatry 2010; 167:998Link, Google Scholar
8 : Copy number variations and risk for schizophrenia in 22q11.2 deletion syndrome. Hum Mol Genet 2008; 17:4045–4053Crossref, Medline, Google Scholar
9 : Whole-genome sequencing suggests schizophrenia risk mechanisms in humans with 22q11.2 deletion syndrome. G3 (Bethesda) 2015; 5:2453–2461Crossref, Medline, Google Scholar
10 : MicroRNA dysregulation, gene networks, and risk for schizophrenia in 22q11.2 deletion syndrome. Front Neurol 2014; 5:238Crossref, Medline, Google Scholar
11 : Explaining the variable penetrance of CNVs: parental intelligence modulates expression of intellectual impairment caused by the 22q11.2 deletion. Am J Med Genet B Neuropsychiatr Genet 2016; 171:790–796Crossref, Medline, Google Scholar
12 : Psychiatric disorders from childhood to adulthood in 22q11.2 deletion syndrome: results from the International Consortium on Brain and Behavior in 22q11.2 Deletion Syndrome. Am J Psychiatry 2014; 171:627–639Link, Google Scholar
13 : Rare copy number variants and congenital heart defects in the 22q11.2 deletion syndrome. Hum Genet 2016; 135:273–285Crossref, Medline, Google Scholar
14 : Pathogenic rare copy number variants in community-based schizophrenia suggest a potential role for clinical microarrays. Hum Mol Genet 2013; 22:4485–4501Crossref, Medline, Google Scholar
15 : Recent genetic findings in schizophrenia and their therapeutic relevance. J Psychopharmacol 2015; 29:85–96Crossref, Medline, Google Scholar
16 : Contribution of copy number variants to schizophrenia from a genome-wide study of 41,321 subjects. Nat Genet 2017; 49:27–35Crossref, Medline, Google Scholar
17 : Integrated genotype calling and association analysis of SNPs, common copy number polymorphisms and rare CNVs. Nat Genet 2008; 40:1253–1260Crossref, Medline, Google Scholar
18 : Comprehensive assessment of array-based platforms and calling algorithms for detection of copy number variants. Nat Biotechnol 2011; 29:512–520Crossref, Medline, Google Scholar
19 : miRBase: integrating microRNA annotation and deep-sequencing data. Nucleic Acids Res 2011; 39:D152–D157Crossref, Medline, Google Scholar
20 : miRNA-mediated risk for schizophrenia in 22q11.2 deletion syndrome. Front Genet 2012; 3:291Crossref, Medline, Google Scholar
21 : De novo CNV analysis implicates specific abnormalities of postsynaptic signalling complexes in the pathogenesis of schizophrenia. Mol Psychiatry 2012; 17:142–153Crossref, Medline, Google Scholar
22 : The GeneMANIA prediction server: biological network integration for gene prioritization and predicting gene function. Nucleic Acids Res 2010; 38:W214–W220Crossref, Medline, Google Scholar
23 : Integration of biological networks and gene expression data using Cytoscape. Nat Protoc 2007; 2:2366–2382Crossref, Medline, Google Scholar
24 : Structures and molecular mechanisms for common 15q13.3 microduplications involving CHRNA7: benign or pathological? Hum Mutat 2010; 31:840–850Crossref, Medline, Google Scholar
25 : Recurrent duplications of 17q12 associated with variable phenotypes. Am J Med Genet A 2015; 167A:3038–3045Crossref, Medline, Google Scholar
26 : 15q13.3 duplication in two patients with childhood-onset schizophrenia. Am J Med Genet B Neuropsychiatr Genet 2016; 171:777–783Crossref, Medline, Google Scholar
27 : Copy number variation in schizophrenia in Sweden. Mol Psychiatry 2014; 19:762–773Crossref, Medline, Google Scholar
28 : Association of common copy number variants at the glutathione S-transferase genes and rare novel genomic changes with schizophrenia. Mol Psychiatry 2010; 15:1023–1033Crossref, Medline, Google Scholar
29 : Copy number variants for schizophrenia and related psychotic disorders in Oceanic Palau: risk and transmission in extended pedigrees. Biol Psychiatry 2011; 70:1115–1121Crossref, Medline, Google Scholar
30 : Strong association of de novo copy number mutations with sporadic schizophrenia. Nat Genet 2008; 40:880–885Crossref, Medline, Google Scholar
31 : A polygenic burden of rare disruptive mutations in schizophrenia. Nature 2014; 506:185–190Crossref, Medline, Google Scholar
32 : Microdeletions of 3q29 confer high risk for schizophrenia. Am J Hum Genet 2010; 87:229–236Crossref, Medline, Google Scholar
33 : Microduplications of 16p11.2 are associated with schizophrenia. Nat Genet 2009; 41:1223–1227Crossref, Medline, Google Scholar
34 : Molecular findings among patients referred for clinical whole-exome sequencing. JAMA 2014; 312:1870–1879Crossref, Medline, Google Scholar
35 : The penetrance of copy number variations for schizophrenia and developmental delay. Biol Psychiatry 2014; 75:378–385Crossref, Medline, Google Scholar
36 : Phenotypic heterogeneity of genomic disorders and rare copy-number variants. N Engl J Med 2012; 367:1321–1331Crossref, Medline, Google Scholar
37 : Schizophrenia two-hit hypothesis in velo-cardio facial syndrome. Am J Med Genet B Neuropsychiatr Genet 2013; 162B:177–182Crossref, Medline, Google Scholar
38 : Patterns of genic intolerance of rare copy number variation in 59,898 human exomes. Nat Genet 2016; 48:1107–1111Crossref, Medline, Google Scholar
39 : De novo variants in sporadic cases of childhood onset schizophrenia. Eur J Hum Genet 2016; 24:944–948Crossref, Medline, Google Scholar
40 : RIM proteins tether Ca2+ channels to presynaptic active zones via a direct PDZ-domain interaction. Cell 2011; 144:282–295Crossref, Medline, Google Scholar
41 : The nogo receptor family restricts synapse number in the developing hippocampus. Neuron 2012; 73:466–481Crossref, Medline, Google Scholar
42 : Rare mutations in N-methyl-d-aspartate glutamate receptors in autism spectrum disorders and schizophrenia. Transl Psychiatry 2011; 1:e55Crossref, Medline, Google Scholar
43 : Gene expression elucidates functional impact of polygenic risk for schizophrenia. Nat Neurosci 2016; 19:1442–1453Crossref, Medline, Google Scholar
44 : Practical guidelines for managing adults with 22q11.2 deletion syndrome. Genet Med 2015; 17:599–609Crossref, Medline, Google Scholar
45 : A higher mutational burden in females supports a “female protective model” in neurodevelopmental disorders. Am J Hum Genet 2014; 94:415–425Crossref, Medline, Google Scholar



